Adapted by Nelson Nuñez-Rodriguez
Conditions of Use:

Unless otherwise noted, this work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License.
Chapters derived from:
By David W. Ball, John W. Hill, and Rhonda J. Scott
 Attribution-NonCommercial-ShareAlike
Attribution-NonCommercial-ShareAlike
CC BY-NC-SA
Click on the printer icon at the bottom of the screen
![]()
Make sure that your printout includes all content from the page. If it doesn't, try opening this guide in a different browser and printing from there (sometimes Internet Explorer works better, sometimes Chrome, sometimes Firefox, etc.).
If the above process produces printouts with errors or overlapping text or images, try this method:
Each of the thousands of naturally occurring proteins has its own characteristic amino acid composition and sequence that result in a unique three-dimensional shape. Since the 1950s, scientists have determined the amino acid sequences and three-dimensional conformation of numerous proteins and thus obtained important clues on how each protein performs its specific function in the body.
Proteins are compounds of high molar mass consisting largely or entirely of chains of amino acids. Because of their great complexity, protein molecules cannot be classified on the basis of specific structural similarities, as carbohydrates and lipids are categorized. The two major structural classifications of proteins are based on far more general qualities: whether the protein is (1) fiberlike and insoluble or (2) globular and soluble. Some proteins, such as those that compose hair, skin, muscles, and connective tissue, are fiberlike. These fibrous proteins are insoluble in water and usually serve structural, connective, and protective functions. Examples of fibrous proteins are keratins, collagens, myosins, and elastins. Hair and the outer layer of skin are composed of keratin. Connective tissues contain collagen. Myosins are muscle proteins and are capable of contraction and extension. Elastins are found in ligaments and the elastic tissue of artery walls.
Globular proteins, the other major class, are soluble in aqueous media. In these proteins, the chains are folded so that the molecule as a whole is roughly spherical. Familiar examples include egg albumin from egg whites and serum albumin in blood. Serum albumin plays a major role in transporting fatty acids and maintaining a proper balance of osmotic pressures in the body. Hemoglobin and myoglobin, which are important for binding oxygen, are also globular proteins.
The structure of proteins is generally described as having four organizational levels. The first of these is the primary structure, which is the number and sequence of amino acids in a protein’s polypeptide chain or chains, beginning with the free amino group and maintained by the peptide bonds connecting each amino acid to the next. The primary structure of insulin, composed of 51 amino acids, is shown in Figure 9.1 "Primary Structure of Human Insulin".
Figure 9.1 Primary Structure of Human Insulin

Human insulin, whose amino acid sequence is shown here, is a hormone that is required for the proper metabolism of glucose.
A protein molecule is not a random tangle of polypeptide chains. Instead, the chains are arranged in unique but specific conformations. The term secondary structure refers to the fixed arrangement of the polypeptide backbone. On the basis of X ray studies, Linus Pauling and Robert Corey postulated that certain proteins or portions of proteins twist into a spiral or a helix. This helix is stabilized by intrachain hydrogen bonding between the carbonyl oxygen atom of one amino acid and the amide hydrogen atom four amino acids up the chain (located on the next turn of the helix) and is known as a right-handed α-helix. X ray data indicate that this helix makes one turn for every 3.6 amino acids, and the side chains of these amino acids project outward from the coiled backbone (Figure 9.2 "A Ball-and-Stick Model of an α-Helix"). The α-keratins, found in hair and wool, are exclusively α-helical in conformation. Some proteins, such as gamma globulin, chymotrypsin, and cytochrome c, have little or no helical structure. Others, such as hemoglobin and myoglobin, are helical in certain regions but not in others.
Figure 9.2 A Ball-and-Stick Model of an α-Helix

This ball-and-stick model shows the intrachain hydrogen bonding between carbonyl oxygen atoms and amide hydrogen atoms. Each turn of the helix spans 3.6 amino acids. Note that the side chains (represented as green spheres) point out from the helix.
Another common type of secondary structure, called the β-pleated sheet conformation, is a sheetlike arrangement in which two or more extended polypeptide chains (or separate regions on the same chain) are aligned side by side. The aligned segments can run either parallel or antiparallel—that is, the N-terminals can face in the same direction on adjacent chains or in different directions—and are connected by interchain hydrogen bonding (Figure 9.3 "A Ball-and-Stick Model of the β-Pleated Sheet Structure in Proteins"). The β-pleated sheet is particularly important in structural proteins, such as silk fibroin. It is also seen in portions of many enzymes, such as carboxypeptidase A and lysozyme.
Figure 9.3 A Ball-and-Stick Model of the β-Pleated Sheet Structure in Proteins
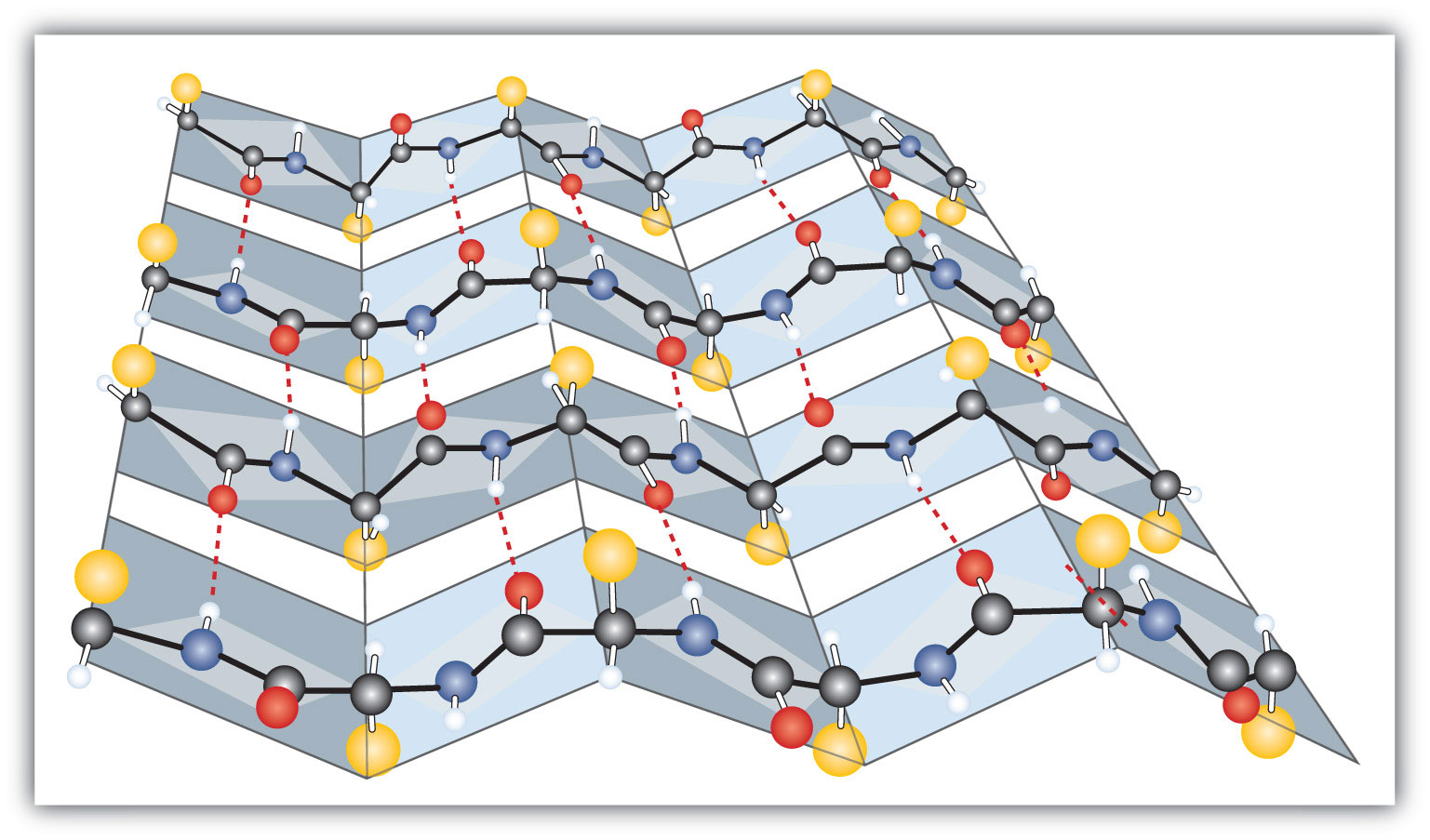
The side chains extend above or below the sheet and alternate along the chain. The protein chains are held together by interchain hydrogen bonding.
Tertiary structure refers to the unique three-dimensional shape of the protein as a whole, which results from the folding and bending of the protein backbone. The tertiary structure is intimately tied to the proper biochemical functioning of the protein. Figure 9.4 "A Ribbon Model of the Three-Dimensional Structure of Insulin" shows a depiction of the three-dimensional structure of insulin.
Figure 9.4 A Ribbon Model of the Three-Dimensional Structure of Insulin
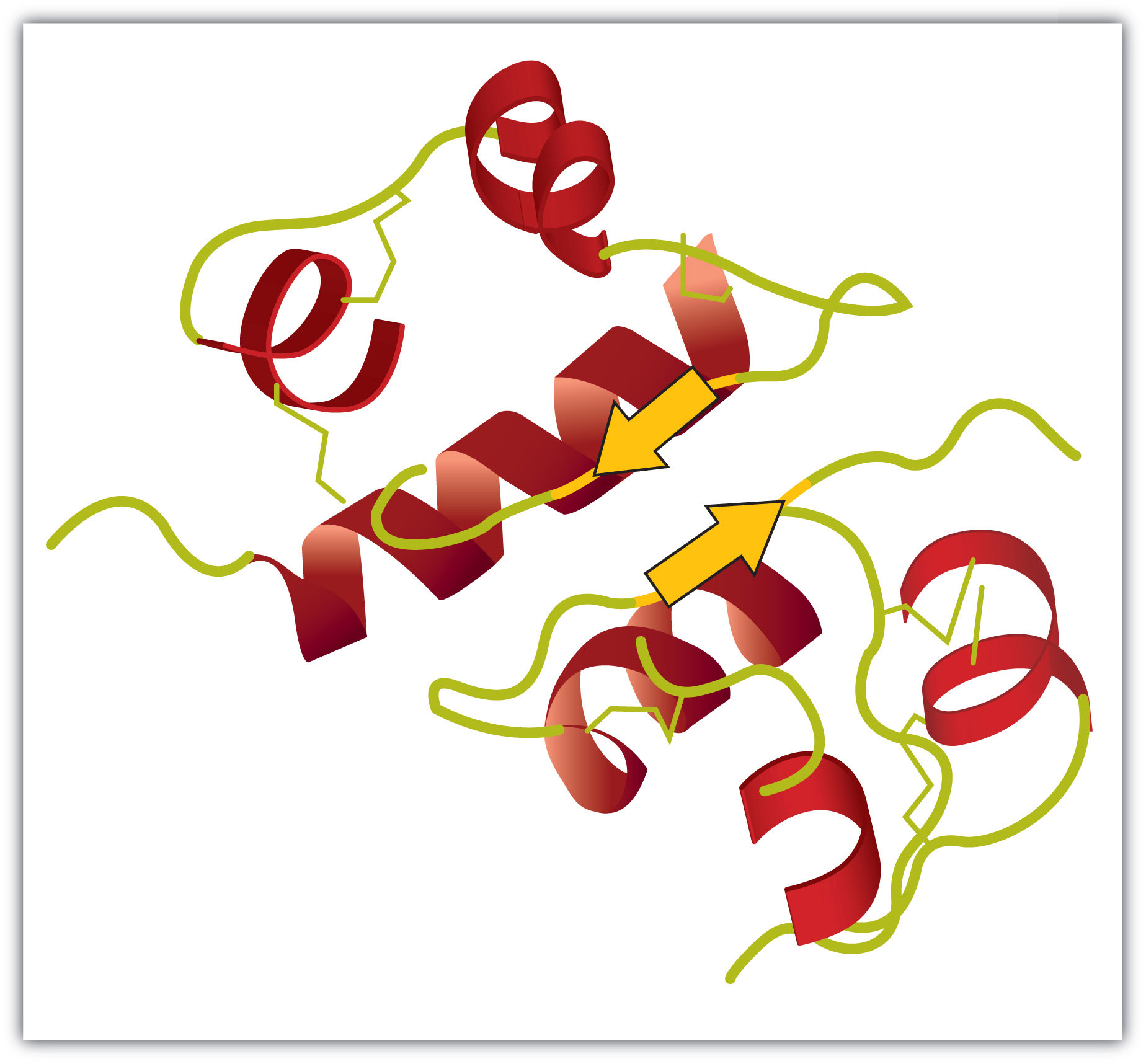
The spiral regions represent sections of the polypeptide chain that have an α-helical structure, while the broad arrows represent β-pleated sheet structures.
Four major types of attractive interactions determine the shape and stability of the tertiary structure of proteins.
1. Ionic bonding. Ionic bonds result from electrostatic attractions between positively and negatively charged side chains of amino acids. For example, the mutual attraction between an aspartic acid carboxylate ion and a lysine ammonium ion helps to maintain a particular folded area of a protein (part (a) of Figure 9.5 "Tertiary Protein Structure Interactions").
2. Hydrogen bonding. Hydrogen bonding forms between a highly electronegative oxygen atom or a nitrogen atom and a hydrogen atom attached to another oxygen atom or a nitrogen atom, such as those found in polar amino acid side chains. Hydrogen bonding (as well as ionic attractions) is extremely important in both the intra- and intermolecular interactions of proteins (part (b) of Figure 9.5 "Tertiary Protein Structure Interactions").
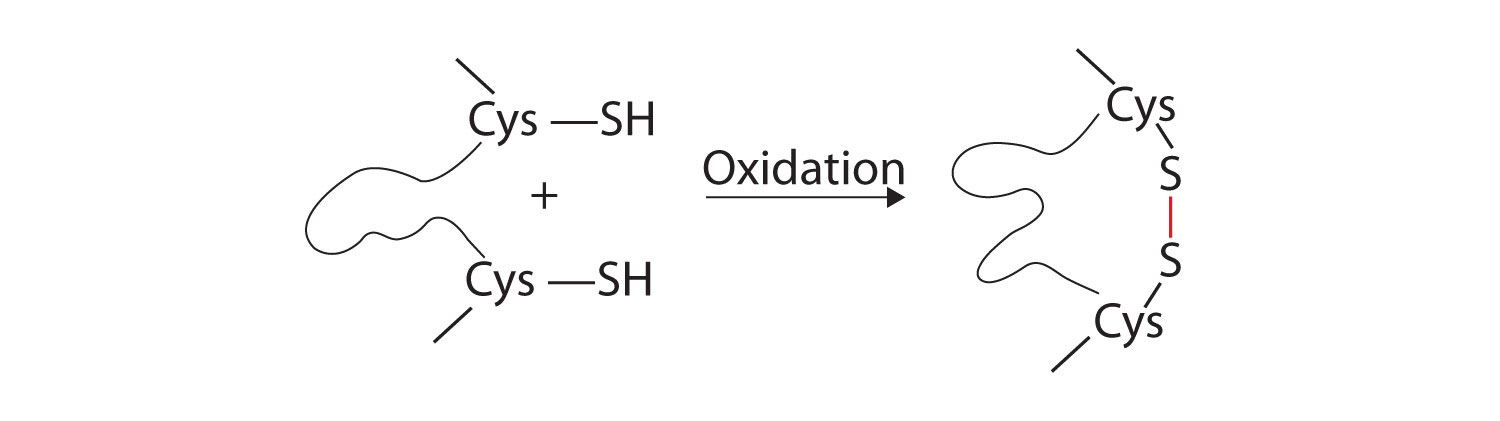
3. Disulfide linkages. Two cysteine amino acid units may be brought close together as the protein molecule folds. Subsequent oxidation and linkage of the sulfur atoms in the highly reactive sulfhydryl (SH) groups leads to the formation of cystine (part (c) of Figure 9.5 "Tertiary Protein Structure Interactions"). Intrachain disulfide linkages are found in many proteins, including insulin (yellow bars in Figure 9.1 "Primary Structure of Human Insulin") and have a strong stabilizing effect on the tertiary structure.
4. Dispersion forces. Dispersion forces arise when a normally nonpolar atom becomes momentarily polar due to an uneven distribution of electrons, leading to an instantaneous dipole that induces a shift of electrons in a neighboring nonpolar atom. Dispersion forces are weak but can be important when other types of interactions are either missing or minimal (part (d) of Figure 9.5 "Tertiary Protein Structure Interactions"). This is the case with fibroin, the major protein in silk, in which a high proportion of amino acids in the protein have nonpolar side chains. The term hydrophobic interaction is often misused as a synonym for dispersion forces. Hydrophobic interactions arise because water molecules engage in hydrogen bonding with other water molecules (or groups in proteins capable of hydrogen bonding). Because nonpolar groups cannot engage in hydrogen bonding, the protein folds in such a way that these groups are buried in the interior part of the protein structure, minimizing their contact with water.
Figure 9.5 Tertiary Protein Structure Interactions
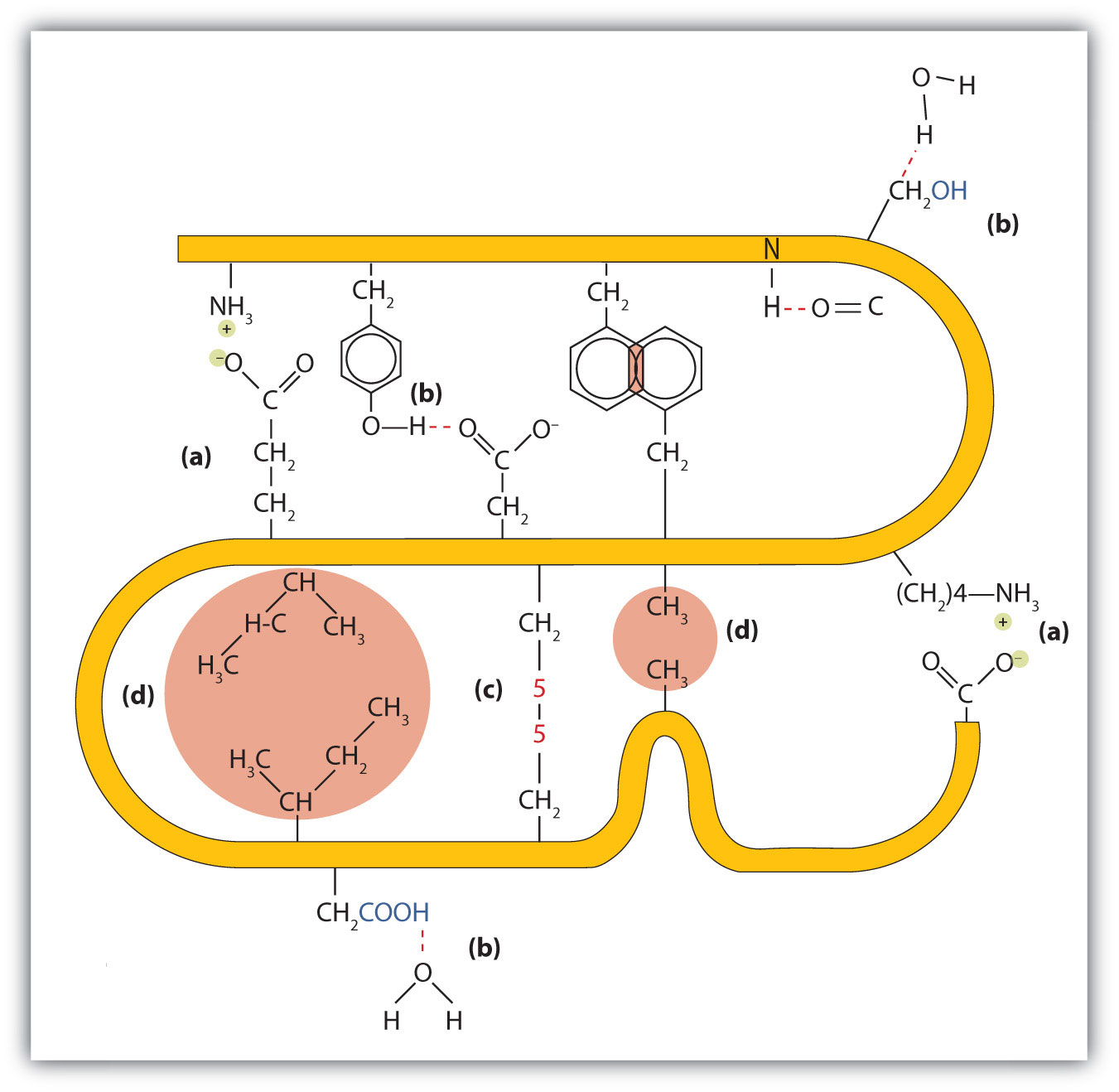
Four interactions stabilize the tertiary structure of a protein: (a) ionic bonding, (b) hydrogen bonding, (c) disulfide linkages, and (d) dispersion forces.
When a protein contains more than one polypeptide chain, each chain is called a subunit. The arrangement of multiple subunits represents a fourth level of structure, the quaternary structure of a protein. Hemoglobin, with four polypeptide chains or subunits, is the most frequently cited example of a protein having quaternary structure (Figure 9.6 "The Quaternary Structure of Hemoglobin"). The quaternary structure of a protein is produced and stabilized by the same kinds of interactions that produce and maintain the tertiary structure. A schematic representation of the four levels of protein structure is in Figure 9.7 "Levels of Structure in Proteins".
Figure 9.6 The Quaternary Structure of Hemoglobin
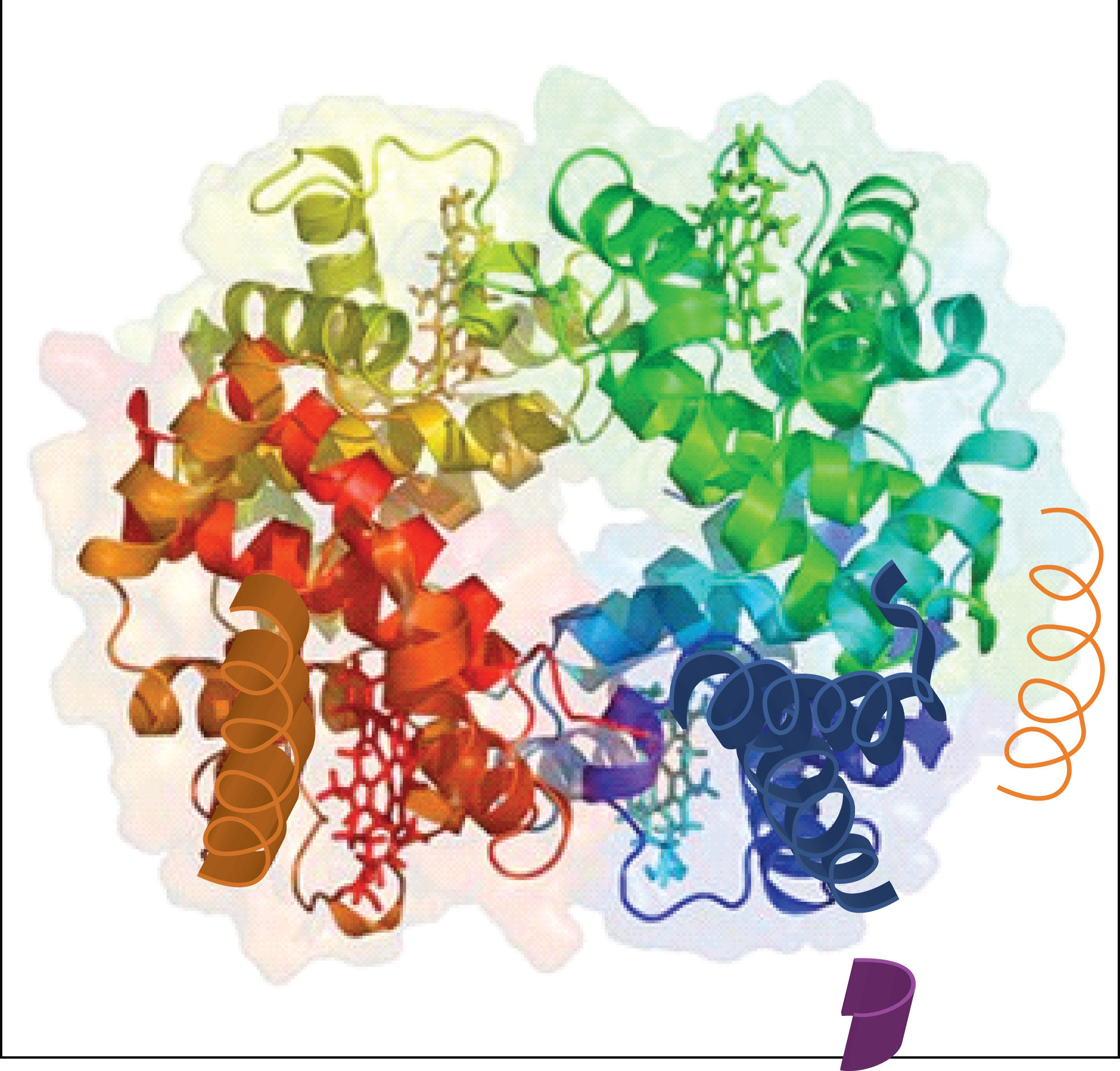
Hemoglobin is a protein that transports oxygen throughout the body.
Source: Image from the RCSB PDB (www.pdb.org) of PDB ID 1I3D (R.D. Kidd, H.M. Baker, A.J. Mathews, T. Brittain, E.N. Baker (2001) Oligomerization and ligand binding in a homotetrameric hemoglobin: two high-resolution crystal structures of hemoglobin Bart's (gamma(4)), a marker for alpha-thalassemia. Protein Sci. 1739–1749).
Figure 9.7 Levels of Structure in Proteins
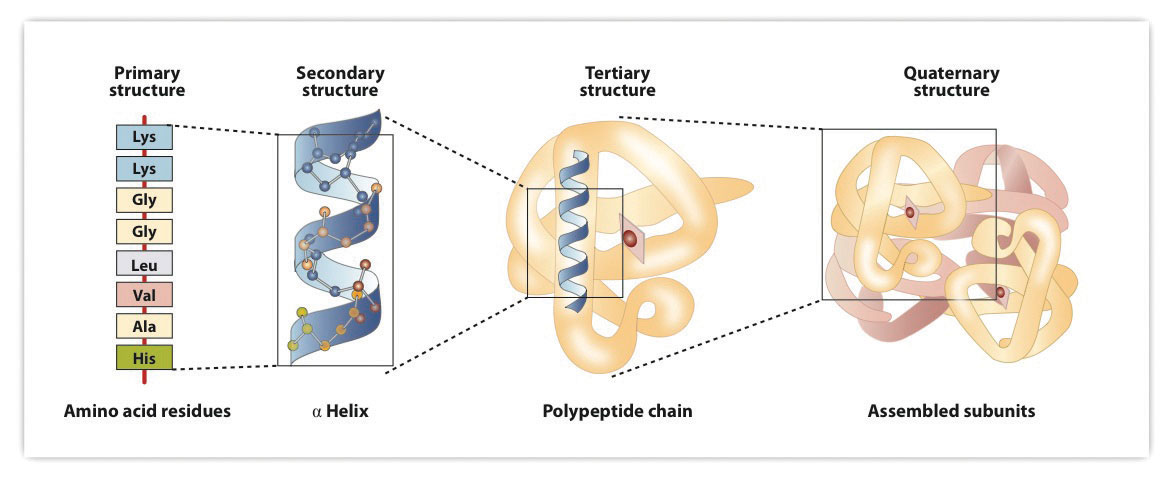
The primary structure consists of the specific amino acid sequence. The resulting peptide chain can twist into an α-helix, which is one type of secondary structure. This helical segment is incorporated into the tertiary structure of the folded polypeptide chain. The single polypeptide chain is a subunit that constitutes the quaternary structure of a protein, such as hemoglobin that has four polypeptide chains.
The highly organized structures of proteins are truly masterworks of chemical architecture. But highly organized structures tend to have a certain delicacy, and this is true of proteins. Denaturation is the term used for any change in the three-dimensional structure of a protein that renders it incapable of performing its assigned function. A denatured protein cannot do its job. (Sometimes denaturation is equated with the precipitation or coagulation of a protein; our definition is a bit broader.) A wide variety of reagents and conditions, such as heat, organic compounds, pH changes, and heavy metal ions can cause protein denaturation (Table 9.1 "Protein Denaturation Methods").
Table 9.1 Protein Denaturation Methods
| Method | Effect on Protein Structure |
|---|---|
| Heat above 50°C or ultraviolet (UV) radiation | Heat or UV radiation supplies kinetic energy to protein molecules, causing their atoms to vibrate more rapidly and disrupting relatively weak hydrogen bonding and dispersion forces. |
| Use of organic compounds, such as ethyl alcohol | These compounds are capable of engaging in intermolecular hydrogen bonding with protein molecules, disrupting intramolecular hydrogen bonding within the protein. |
| Salts of heavy metal ions, such as mercury, silver, and lead | These ions form strong bonds with the carboxylate anions of the acidic amino acids or SH groups of cysteine, disrupting ionic bonds and disulfide linkages. |
| Alkaloid reagents, such as tannic acid (used in tanning leather) | These reagents combine with positively charged amino groups in proteins to disrupt ionic bonds. |
Anyone who has fried an egg has observed denaturation. The clear egg white turns opaque as the albumin denatures and coagulates. No one has yet reversed that process. However, given the proper circumstances and enough time, a protein that has unfolded under sufficiently gentle conditions can refold and may again exhibit biological activity (Figure 9.8 "Denaturation and Renaturation of a Protein"). Such evidence suggests that, at least for these proteins, the primary structure determines the secondary and tertiary structure. A given sequence of amino acids seems to adopt its particular three-dimensional arrangement naturally if conditions are right.
Figure 9.8 Denaturation and Renaturation of a Protein
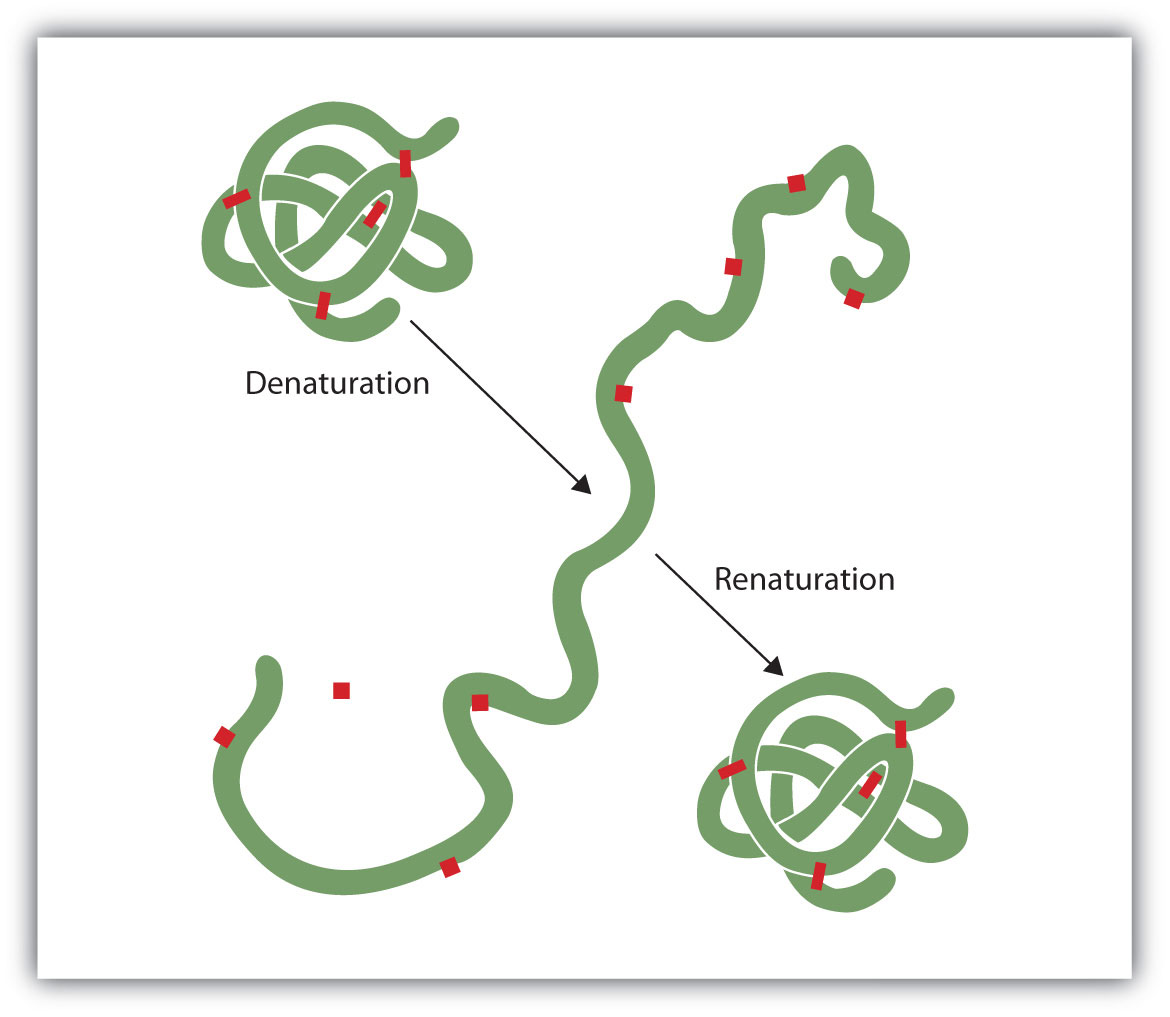
The denaturation (unfolding) and renaturation (refolding) of a protein is depicted. The red boxes represent stabilizing interactions, such as disulfide linkages, hydrogen bonding, and/or ionic bonds.
The primary structures of proteins are quite sturdy. In general, fairly vigorous conditions are needed to hydrolyze peptide bonds. At the secondary through quaternary levels, however, proteins are quite vulnerable to attack, though they vary in their vulnerability to denaturation. The delicately folded globular proteins are much easier to denature than are the tough, fibrous proteins of hair and skin.
What is the predominant attractive force that stabilizes the formation of secondary structure in proteins?
Distinguish between the tertiary and quaternary levels of protein structure.
Briefly describe four ways in which a protein could be denatured.
hydrogen bonding
Tertiary structure refers to the unique three-dimensional shape of a single polypeptide chain, while quaternary structure describes the interaction between multiple polypeptide chains for proteins that have more than one polypeptide chain.
(1) heat a protein above 50°C or expose it to UV radiation; (2) add organic solvents, such as ethyl alcohol, to a protein solution; (3) add salts of heavy metal ions, such as mercury, silver, or lead; and (4) add alkaloid reagents such as tannic acid
1. Classify each protein as fibrous or globular.
a. albumin
b. myosin
c. fibroin
2. Classify each protein as fibrous or globular.
a. hemoglobin
b. keratin
c. myoglobin
3. What name is given to the predominant secondary structure found in silk?
4. What name is given to the predominant secondary structure found in wool protein?
5. A protein has a tertiary structure formed by interactions between the side chains of the following pairs of amino acids. For each pair, identify the strongest type of interaction between these amino acids.
a. aspartic acid and lysine
b. phenylalanine and alanine
c. serine and lysine
d. two cysteines
6. A protein has a tertiary structure formed by interactions between the side chains of the following pairs of amino acids. For each pair, identify the strongest type of interaction between these amino acids.
a. valine and isoleucine
b. asparagine and serine
c. glutamic acid and arginine
d. tryptophan and methionine
7. What level(s) of protein structure is(are) ordinarily disrupted in denaturation? What level(s) is(are) not?
8. Which class of proteins is more easily denatured—fibrous or globular?
1.
a. globular
b. fibrous
c. fibrous
3. β-pleated sheet
5.
a. ionic bonding
b. dispersion forces
c. dispersion forces
d. disulfide linkage
7. Protein denaturation disrupts the secondary, tertiary, and quaternary levels of structure. Only primary structure is unaffected by denaturation.
A catalyst is any substance that increases the rate or speed of a chemical reaction without being changed or consumed in the reaction. Enzymes are biological catalysts, and nearly all of them are proteins. The reaction rates attained by enzymes are truly amazing. In their presence, reactions occur at rates that are a million (106) or more times faster than would be attainable in their absence. What is even more amazing is that enzymes perform this function at body temperature (~37°C) and physiological pH (pH ~7), rather than at the conditions that are typically necessary to increase reaction rates (high temperature or pressure, the use of strong oxidizing or reducing agents or strong acids or bases, or a combination of any of these). In addition, enzymes are highly specific in their action; that is, each enzyme catalyzes only one type of reaction in only one compound or a group of structurally related compounds. The compound or compounds on which an enzyme acts are known as its substrates.
Hundreds of enzymes have been purified and studied in an effort to understand how they work so effectively and with such specificity. The resulting knowledge has been used to design drugs that inhibit or activate particular enzymes. An example is the intensive research to improve the treatment of or find a cure for acquired immunodeficiency syndrome (AIDS). AIDS is caused by the human immunodeficiency virus (HIV). Researchers are studying the enzymes produced by this virus and are developing drugs intended to block the action of those enzymes without interfering with enzymes produced by the human body. Several of these drugs have now been approved for use by AIDS patients.
The first enzymes to be discovered were named according to their source or method of discovery. The enzyme pepsin, which aids in the hydrolysis of proteins, is found in the digestive juices of the stomach (Greek pepsis, meaning “digestion”). Papain, another enzyme that hydrolyzes protein (in fact, it is used in meat tenderizers), is isolated from papayas. As more enzymes were discovered, chemists recognized the need for a more systematic and chemically informative identification scheme.
In the current numbering and naming scheme, under the oversight of the Nomenclature Commission of the International Union of Biochemistry, enzymes are arranged into six groups according to the general type of reaction they catalyze (Table 9.2 "Classes of Enzymes"), with subgroups and secondary subgroups that specify the reaction more precisely. Each enzyme is assigned a four-digit number, preceded by the prefix EC—for enzyme classification—that indicates its group, subgroup, and so forth. This is demonstrated in Table 9.3 "Assignment of an Enzyme Classification Number" for alcohol dehydrogenase. Each enzyme is also given a name consisting of the root of the name of its substrate or substrates and the -ase suffix. Thus urease is the enzyme that catalyzes the hydrolysis of urea.
Table 9.2 Classes of Enzymes
| Class | Type of Reaction Catalyzed | Examples |
|---|---|---|
| oxidoreductases | oxidation-reduction reactions | Dehydrogenases catalyze oxidation-reduction reactions involving hydrogen and reductases catalyze reactions in which a substrate is reduced. |
| transferases | transfer reactions of groups, such as methyl, amino, and acetyl | Transaminases catalyze the transfer of amino group, and kinases catalyze the transfer of a phosphate group. |
| hydrolases | hydrolysis reactions | Lipases catalyze the hydrolysis of lipids, and proteases catalyze the hydrolysis of proteins |
| lyases | reactions in which groups are removed without hydrolysis or addition of groups to a double bond | Decarboxylases catalyze the removal of carboxyl groups. |
| isomerases | reactions in which a compound is converted to its isomer | Isomerases may catalyze the conversion of an aldose to a ketose, and mutases catalyze reactions in which a functional group is transferred from one atom in a substrate to another. |
| ligases | reactions in which new bonds are formed between carbon and another atom; energy is required | Synthetases catalyze reactions in which two smaller molecules are linked to form a larger one. |
Table 9.3 Assignment of an Enzyme Classification Number
| Alcohol Dehydrogenase: EC 1.1.1.1 |
|---|
| The first digit indicates that this enzyme is an oxidoreductase; that is, an enzyme that catalyzes an oxidation-reduction reaction. |
| The second digit indicates that this oxidoreductase catalyzes a reaction involving a primary or secondary alcohol. |
| The third digit indicates that either the coenzyme NAD+ or NADP+ is required for this reaction. |
| The fourth digit indicates that this was the first enzyme isolated, characterized, and named using this system of nomenclature. |
| The systematic name for this enzyme is alcohol:NAD+ oxidoreductase, while the recommended or common name is alcohol dehydrogenase. |
Reaction catalyzed: 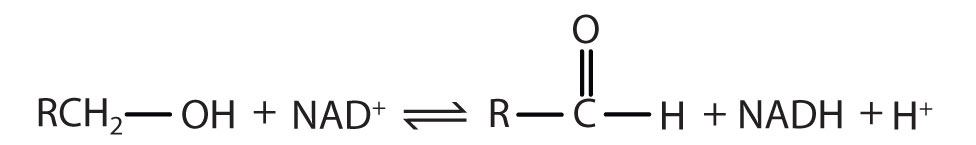 |
In the small intestine, sucrose is hydrolyzed to form glucose and fructose in a reaction catalyzed by sucrase.
Identify the substrate in this reaction.
Name the enzyme.
sucrose
sucrase
1. Identify the substrate catalyzed by each enzyme.
a. lactase
b. cellulase
c. peptidase
2. Identify the substrate catalyzed by each enzyme.
a. lipase
b. amylase
c. maltase
3. Identify each type of enzyme.
a. decarboxylase
b. protease
c. transaminase
4. Identify each type of enzyme.
a. dehydrogenase
b. isomerase
c. lipase
1.
a. lactose
b. cellulose
c. peptides
3.
a. lyase
b. hydrolase
c. transferase
Enzyme-catalyzed reactions occur in at least two steps. In the first step, an enzyme molecule (E) and the substrate molecule or molecules (S) collide and react to form an intermediate compound called the enzyme-substrate (E–S) complex. (This step is reversible because the complex can break apart into the original substrate or substrates and the free enzyme.) Once the E–S complex forms, the enzyme is able to catalyze the formation of product (P), which is then released from the enzyme surface:
S + E → E–S E–S → P + E
Hydrogen bonding and other electrostatic interactions hold the enzyme and substrate together in the complex. The structural features or functional groups on the enzyme that participate in these interactions are located in a cleft or pocket on the enzyme surface. This pocket, where the enzyme combines with the substrate and transforms the substrate to product is called the active site of the enzyme (Figure 9.9 "Substrate Binding to the Active Site of an Enzyme"). It possesses a unique conformation (including correctly positioned bonding groups) that is complementary to the structure of the substrate, so that the enzyme and substrate molecules fit together in much the same manner as a key fits into a tumbler lock. In fact, an early model describing the formation of the enzyme-substrate complex was called the lock-and-key model (Figure 9.10 "The Lock-and-Key Model of Enzyme Action"). This model portrayed the enzyme as conformationally rigid and able to bond only to substrates that exactly fit the active site.
Figure 9.9 Substrate Binding to the Active Site of an Enzyme
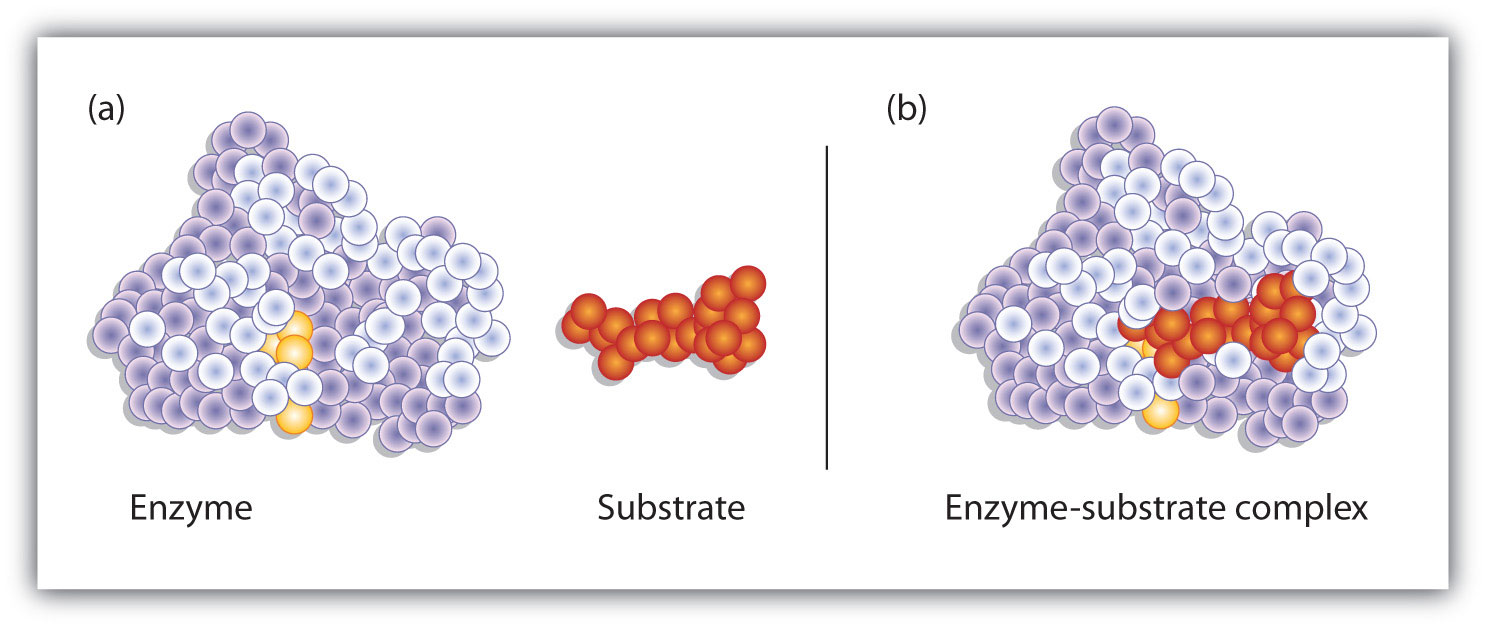
The enzyme dihydrofolate reductase is shown with one of its substrates: NADP+ (a) unbound and (b) bound. The NADP+ (shown in red) binds to a pocket that is complementary to it in shape and ionic properties.
Figure 9.10 The Lock-and-Key Model of Enzyme Action
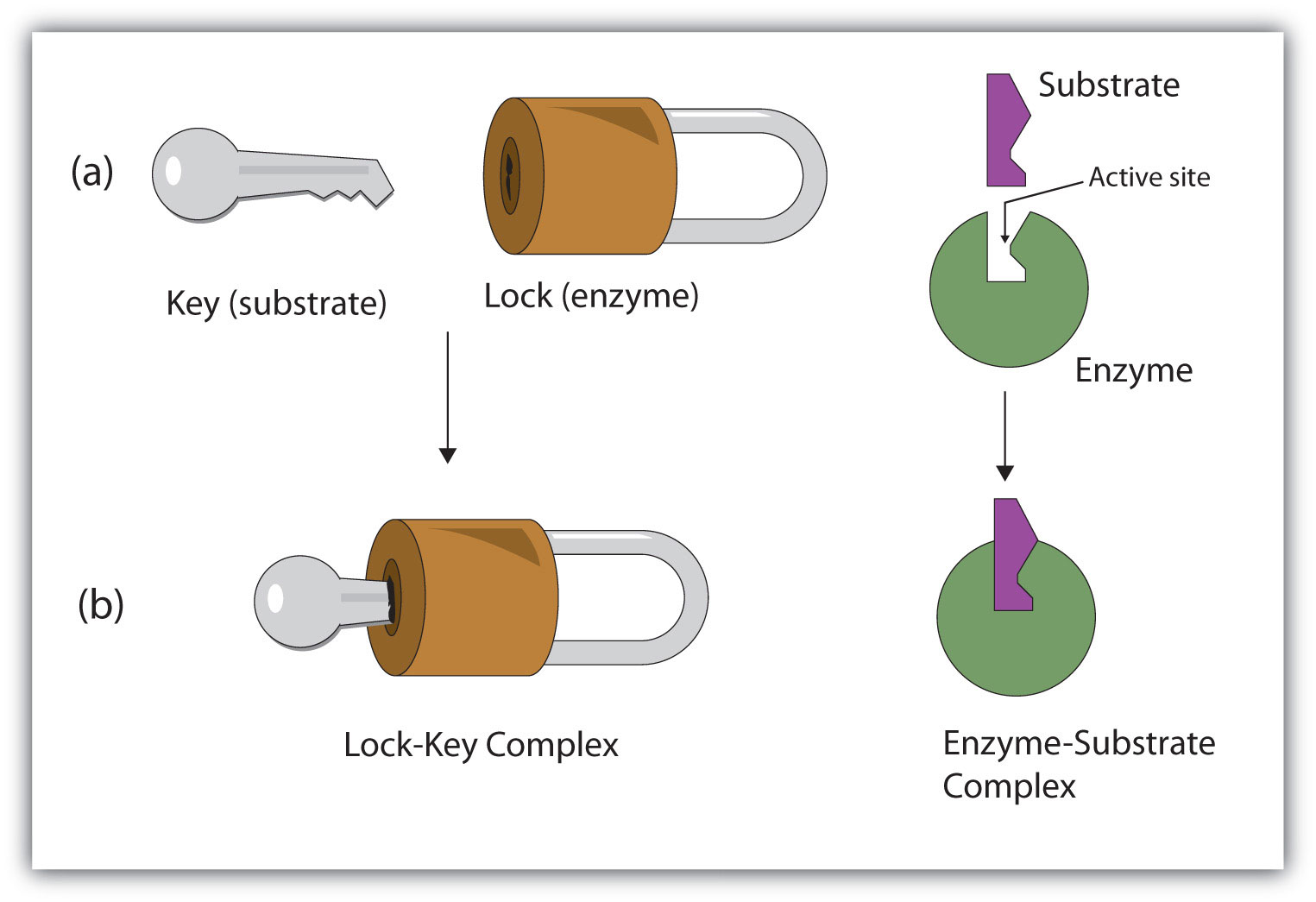
(a) Because the substrate and the active site of the enzyme have complementary structures and bonding groups, they fit together as a key fits a lock. (b) The catalytic reaction occurs while the two are bonded together in the enzyme-substrate complex.
Working out the precise three-dimensional structures of numerous enzymes has enabled chemists to refine the original lock-and-key model of enzyme actions. They discovered that the binding of a substrate often leads to a large conformational change in the enzyme, as well as to changes in the structure of the substrate or substrates. The current theory, known as the induced-fit model, says that enzymes can undergo a change in conformation when they bind substrate molecules, and the active site has a shape complementary to that of the substrate only after the substrate is bound, as shown for hexokinase in Figure 9.11 "The Induced-Fit Model of Enzyme Action". After catalysis, the enzyme resumes its original structure.
Figure 9.11 The Induced-Fit Model of Enzyme Action
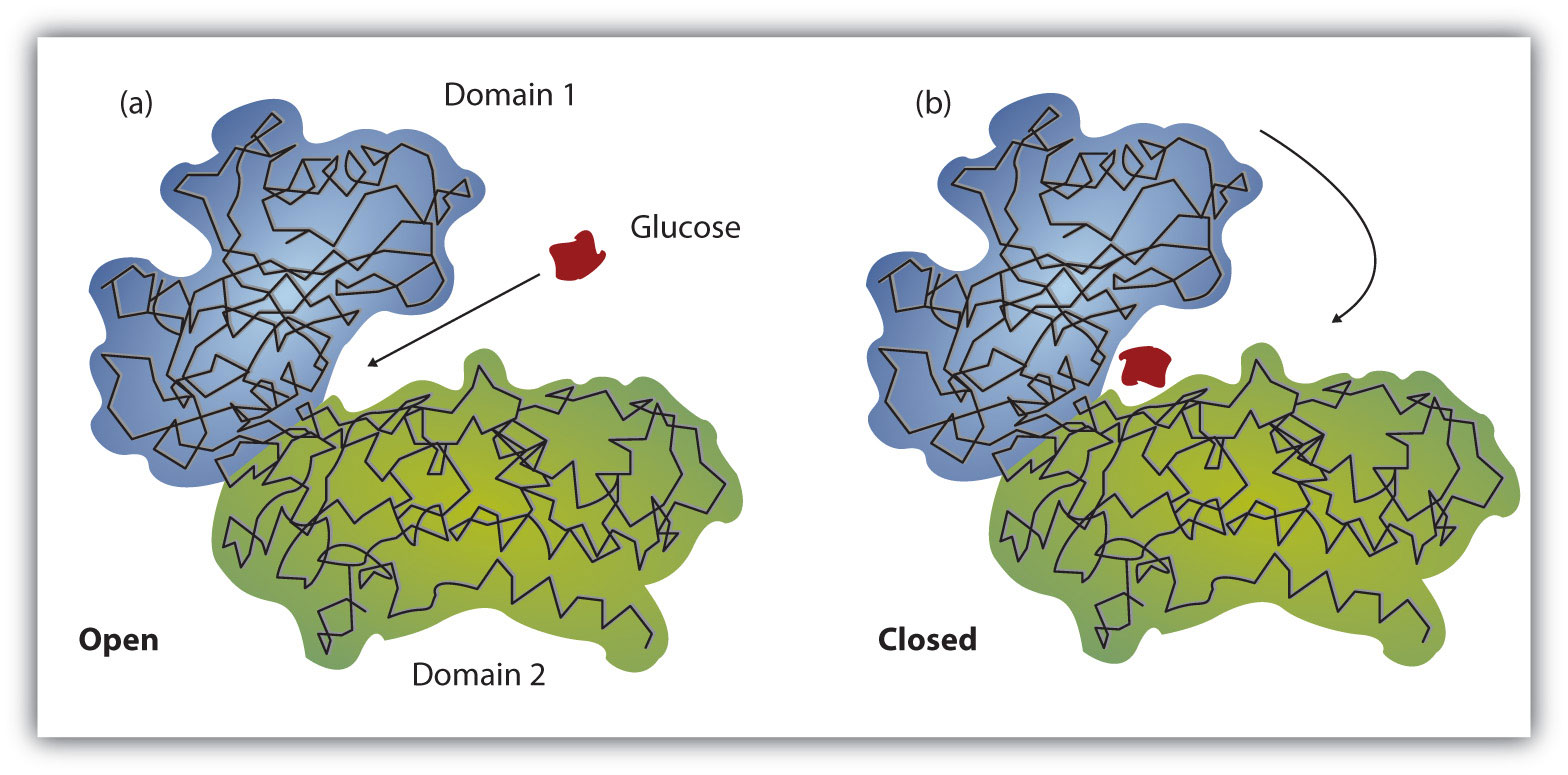
(a) The enzyme hexokinase without its substrate (glucose, shown in red) is bound to the active site. (b) The enzyme conformation changes dramatically when the substrate binds to it, resulting in additional interactions between hexokinase and glucose.
The structural changes that occur when an enzyme and a substrate join together bring specific parts of a substrate into alignment with specific parts of the enzyme’s active site. Amino acid side chains in or near the binding site can then act as acid or base catalysts, provide binding sites for the transfer of functional groups from one substrate to another or aid in the rearrangement of a substrate. The participating amino acids, which are usually widely separated in the primary sequence of the protein, are brought close together in the active site as a result of the folding and bending of the polypeptide chain or chains when the protein acquires its tertiary and quaternary structure. Binding to enzymes brings reactants close to each other and aligns them properly, which has the same effect as increasing the concentration of the reacting compounds.
Solution
What type of interaction would occur between an COO− group present on a substrate molecule and a functional group in the active site of an enzyme?
Suggest an amino acid whose side chain might be in the active site of an enzyme and form the type of interaction you just identified.
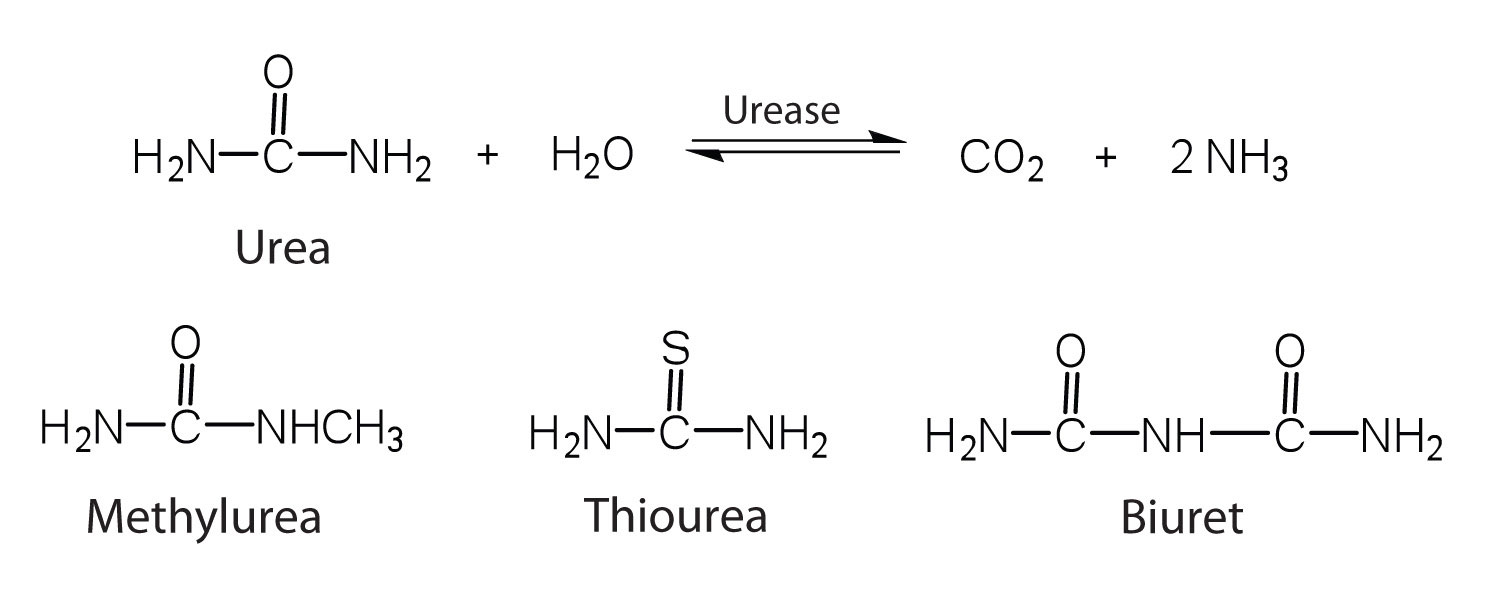
One characteristic that distinguishes an enzyme from all other types of catalysts is its substrate specificity. An inorganic acid such as sulfuric acid can be used to increase the reaction rates of many different reactions, such as the hydrolysis of disaccharides, polysaccharides, lipids, and proteins, with complete impartiality. In contrast, enzymes are much more specific. Some enzymes act on a single substrate, while other enzymes act on any of a group of related molecules containing a similar functional group or chemical bond. Some enzymes even distinguish between D- and L-stereoisomers, binding one stereoisomer but not the other. Urease, for example, is an enzyme that catalyzes the hydrolysis of a single substrate—urea—but not the closely related compounds methyl urea, thiourea, or biuret. The enzyme carboxypeptidase, on the other hand, is far less specific. It catalyzes the removal of nearly any amino acid from the carboxyl end of any peptide or protein.
Enzyme specificity results from the uniqueness of the active site in each different enzyme because of the identity, charge, and spatial orientation of the functional groups located there. It regulates cell chemistry so that the proper reactions occur in the proper place at the proper time. Clearly, it is crucial to the proper functioning of the living cell.
Distinguish between the lock-and-key model and induced-fit model of enzyme action.
Which enzyme has greater specificity—urease or carboxypeptidase? Explain.
The lock-and-key model portrays an enzyme as conformationally rigid and able to bond only to substrates that exactly fit the active site. The induced fit model portrays the enzyme structure as more flexible and is complementary to the substrate only after the substrate is bound.
Urease has the greater specificity because it can bind only to a single substrate. Carboxypeptidase, on the other hand, can catalyze the removal of nearly any amino acid from the carboxyl end of a peptide or protein.
1. What type of interaction would occur between each group present on a substrate molecule and a functional group of the active site in an enzyme?
a. COOH
b. NH3+
c. OH
d. CH(CH3)2
2. What type of interaction would occur between each group present on a substrate molecule and a functional group of the active site in an enzyme?
a. SH
b. NH2
c. C6H5
d. COO−
3. For each functional group in Exercise 1, suggest an amino acid whose side chain might be in the active site of an enzyme and form the type of interaction you identified.
4. For each functional group in Exercise 2, suggest an amino acid whose side chain might be in the active site of an enzyme and form the type of interaction you identified.
1.
a. hydrogen bonding
b. ionic bonding
c. hydrogen bonding
d. dispersion forces
3.
a. The amino acid has a polar side chain capable of engaging in hydrogen bonding; serine (answers will vary).
b. The amino acid has a negatively charged side chain; aspartic acid (answers will vary).
c. The amino acid has a polar side chain capable of engaging in hydrogen bonding; asparagine (answers will vary).
d. The amino acid has a nonpolar side chain; isoleucine (answers will vary).
The single most important property of enzymes is the ability to increase the rates of reactions occurring in living organisms, a property known as catalytic activity. Because most enzymes are proteins, their activity is affected by factors that disrupt protein structure, as well as by factors that affect catalysts in general. Factors that disrupt protein structure, as we saw in Section 9.1 "Proteins", include temperature and pH; factors that affect catalysts in general include reactant or substrate concentration and catalyst or enzyme concentration. The activity of an enzyme can be measured by monitoring either the rate at which a substrate disappears or the rate at which a product forms.
In the presence of a given amount of enzyme, the rate of an enzymatic reaction increases as the substrate concentration increases until a limiting rate is reached, after which further increase in the substrate concentration produces no significant change in the reaction rate (part (a) of Figure 9.12 "Concentration versus Reaction Rate"). At this point, so much substrate is present that essentially all of the enzyme active sites have substrate bound to them. In other words, the enzyme molecules are saturated with substrate. The excess substrate molecules cannot react until the substrate already bound to the enzymes has reacted and been released (or been released without reacting).
Figure 9.12 Concentration versus Reaction Rate
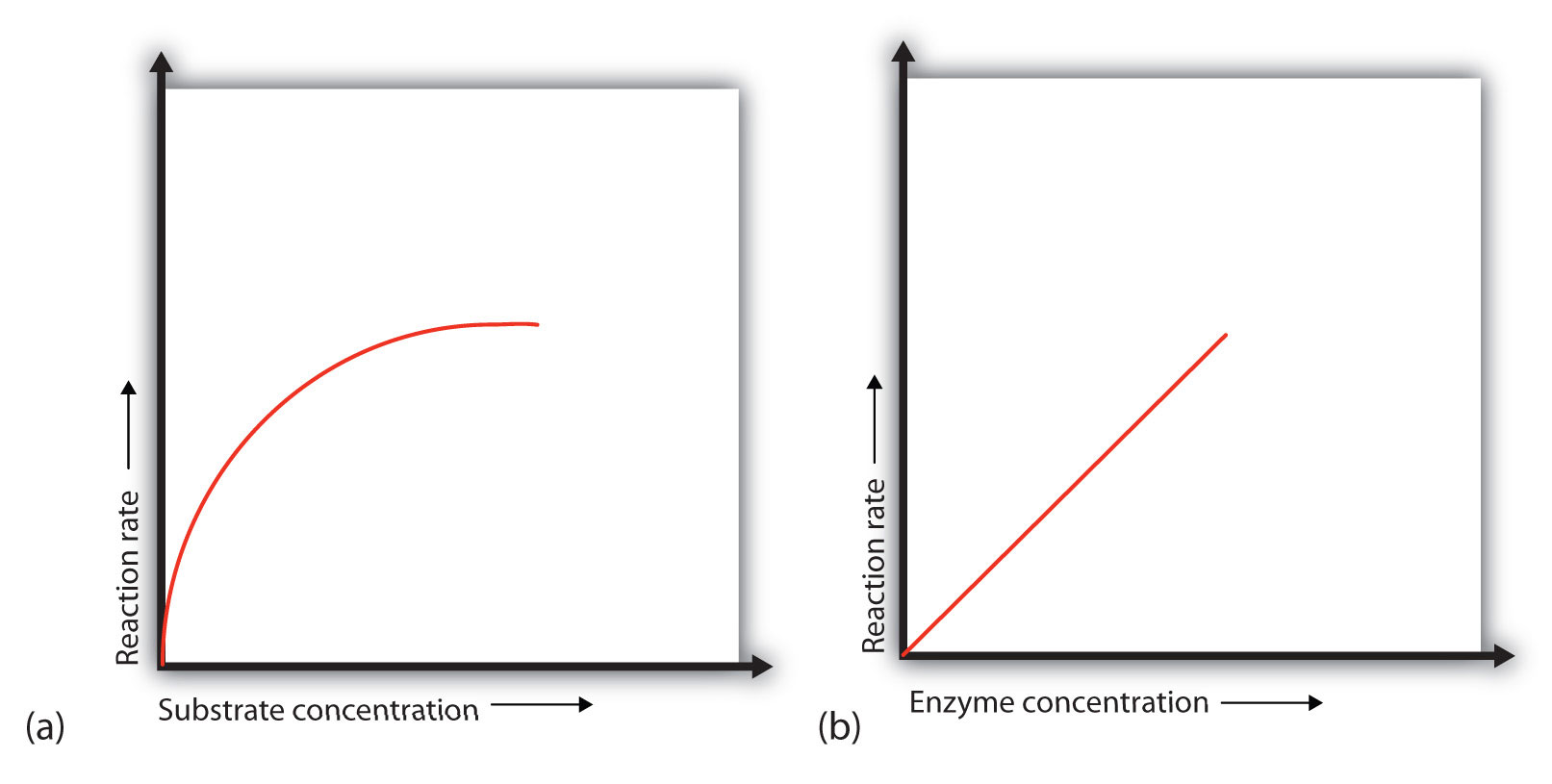
(a) This graph shows the effect of substrate concentration on the rate of a reaction that is catalyzed by a fixed amount of enzyme. (b) This graph shows the effect of enzyme concentration on the reaction rate at a constant level of substrate.
Let’s consider an analogy. Ten taxis (enzyme molecules) are waiting at a taxi stand to take people (substrate) on a 10-minute trip to a concert hall, one passenger at a time. If only 5 people are present at the stand, the rate of their arrival at the concert hall is 5 people in 10 minutes. If the number of people at the stand is increased to 10, the rate increases to 10 arrivals in 10 minutes. With 20 people at the stand, the rate would still be 10 arrivals in 10 minutes. The taxis have been “saturated.” If the taxis could carry 2 or 3 passengers each, the same principle would apply. The rate would simply be higher (20 or 30 people in 10 minutes) before it leveled off.
When the concentration of the enzyme is significantly lower than the concentration of the substrate (as when the number of taxis is far lower than the number of waiting passengers), the rate of an enzyme-catalyzed reaction is directly dependent on the enzyme concentration (part (b) of Figure 9.12 "Concentration versus Reaction Rate"). This is true for any catalyst; the reaction rate increases as the concentration of the catalyst is increased.
A general rule of thumb for most chemical reactions is that a temperature rise of 10°C approximately doubles the reaction rate. To some extent, this rule holds for all enzymatic reactions. After a certain point, however, an increase in temperature causes a decrease in the reaction rate, due to denaturation of the protein structure and disruption of the active site (part (a) of Figure 9.13 "Temperature and pH versus Concentration"). For many proteins, denaturation occurs between 45°C and 55°C. Furthermore, even though an enzyme may appear to have a maximum reaction rate between 40°C and 50°C, most biochemical reactions are carried out at lower temperatures because enzymes are not stable at these higher temperatures and will denature after a few minutes.
Figure 9.13 Temperature and pH versus Concentration
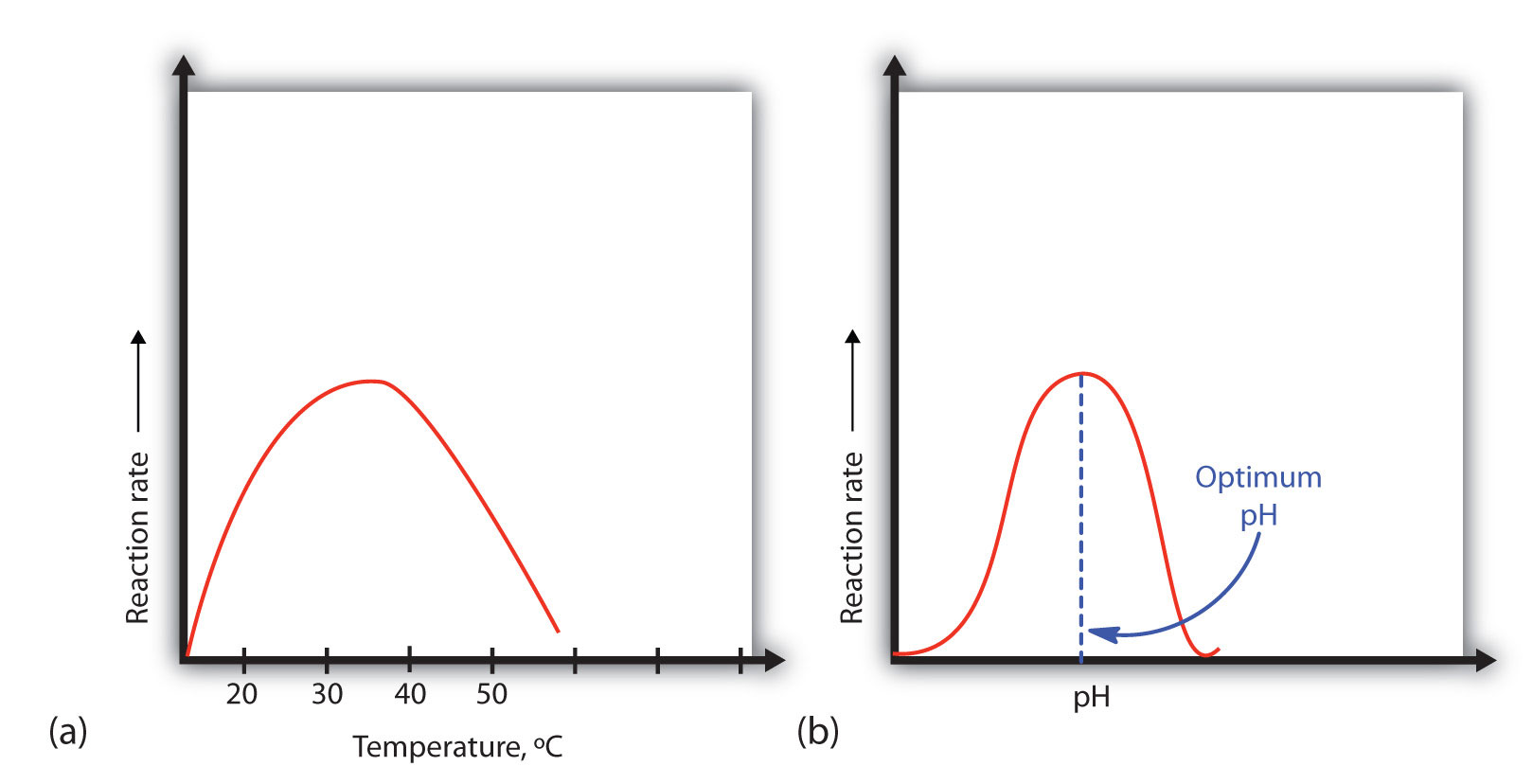
(a) This graph depicts the effect of temperature on the rate of a reaction that is catalyzed by a fixed amount of enzyme. (b) This graph depicts the effect of pH on the rate of a reaction that is catalyzed by a fixed amount of enzyme.
At 0°C and 100°C, the rate of enzyme-catalyzed reactions is nearly zero. This fact has several practical applications. We sterilize objects by placing them in boiling water, which denatures the enzymes of any bacteria that may be in or on them. We preserve our food by refrigerating or freezing it, which slows enzyme activity. When animals go into hibernation in winter, their body temperature drops, decreasing the rates of their metabolic processes to levels that can be maintained by the amount of energy stored in the fat reserves in the animals’ tissues.
Because most enzymes are proteins, they are sensitive to changes in the hydrogen ion concentration or pH. Enzymes may be denatured by extreme levels of hydrogen ions (whether high or low); any change in pH, even a small one, alters the degree of ionization of an enzyme’s acidic and basic side groups and the substrate components as well. Ionizable side groups located in the active site must have a certain charge for the enzyme to bind its substrate. Neutralization of even one of these charges alters an enzyme’s catalytic activity.
An enzyme exhibits maximum activity over the narrow pH range in which a molecule exists in its properly charged form. The median value of this pH range is called the optimum pH of the enzyme (part (b) of Figure 9.13 "Temperature and pH versus Concentration"). With the notable exception of gastric juice (the fluids secreted in the stomach), most body fluids have pH values between 6 and 8. Not surprisingly, most enzymes exhibit optimal activity in this pH range. However, a few enzymes have optimum pH values outside this range. For example, the optimum pH for pepsin, an enzyme that is active in the stomach, is 2.0.
The concentration of substrate X is low. What happens to the rate of the enzyme-catalyzed reaction if the concentration of X is doubled?
What effect does an increase in the enzyme concentration have on the rate of an enzyme-catalyzed reaction?
If the concentration of the substrate is low, increasing its concentration will increase the rate of the reaction.
An increase in the amount of enzyme will increase the rate of the reaction (provided sufficient substrate is present).
1. In non-enzyme-catalyzed reactions, the reaction rate increases as the concentration of reactant is increased. In an enzyme-catalyzed reaction, the reaction rate initially increases as the substrate concentration is increased but then begins to level off, so that the increase in reaction rate becomes less and less as the substrate concentration increases. Explain this difference.
2. Why do enzymes become inactive at very high temperatures?
3. An enzyme has an optimum pH of 7.4. What is most likely to happen to the activity of the enzyme if the pH drops to 6.3? Explain.
4. An enzyme has an optimum pH of 7.2. What is most likely to happen to the activity of the enzyme if the pH increases to 8.5? Explain.
1. In an enzyme-catalyzed reaction, the substrate binds to the enzyme to form an enzyme-substrate complex. If more substrate is present than enzyme, all of the enzyme binding sites will have substrate bound, and further increases in substrate concentration cannot increase the rate.
3. The activity will decrease; a pH of 6.3 is more acidic than 7.4, and one or more key groups in the active site may bind a hydrogen ion, changing the charge on that group.
In Section 9.4 "Enzyme Activity", we noted that enzymes are inactivated at high temperatures and by changes in pH. These are nonspecific factors that would inactivate any enzyme. The activity of enzymes can also be regulated by more specific inhibitors. Many compounds are poisons because they bind covalently to particular enzymes or kinds of enzymes and inactivate them (Table 9.4 "Poisons as Enzyme Inhibitors").
Table 9.4 Poisons as Enzyme Inhibitors
| Poison | Formula | Example of Enzyme Inhibited | Action |
|---|---|---|---|
| arsenate | AsO43− | glyceraldehyde 3-phosphate dehydrogenase | substitutes for phosphate |
| iodoacetate | ICH2COO− | triose phosphate dehydrogenase | binds to cysteine SH group |
| diisopropylfluoro-phosphate (DIFP; a nerve poison) | 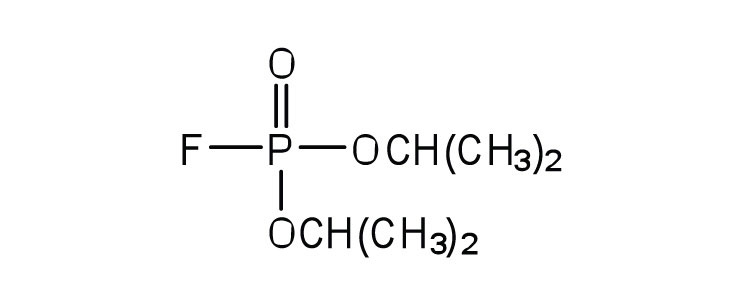 |
acetylcholinesterase | binds to serine OH group |
An irreversible inhibitor inactivates an enzyme by bonding covalently to a particular group at the active site. The inhibitor-enzyme bond is so strong that the inhibition cannot be reversed by the addition of excess substrate. The nerve gases, especially DIFP, irreversibly inhibit biological systems by forming an enzyme-inhibitor complex with a specific OH group of serine situated at the active sites of certain enzymes. The peptidases trypsin and chymotrypsin contain serine groups at the active site and are inhibited by DIFP.
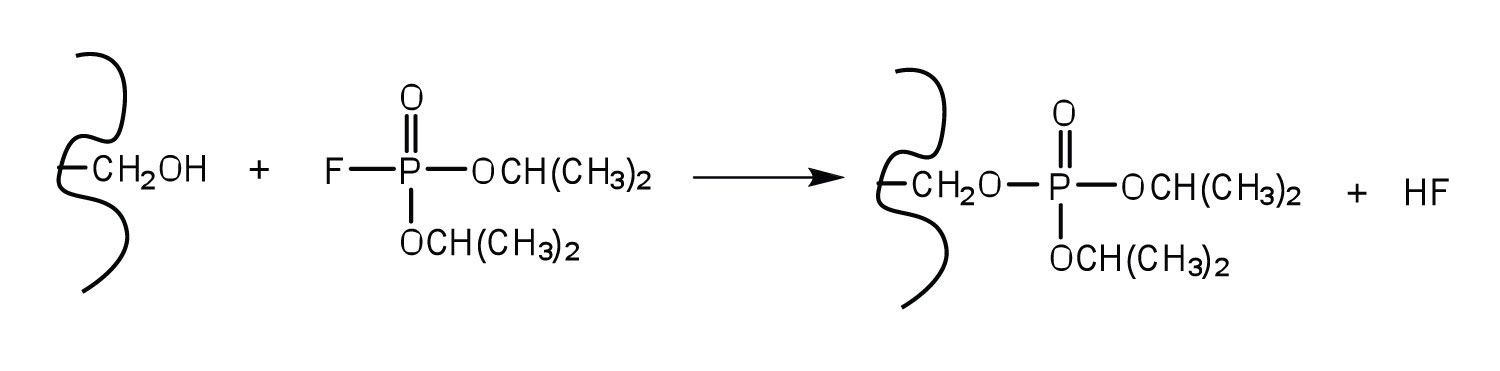
A reversible inhibitor inactivates an enzyme through noncovalent, more easily reversed, interactions. Unlike an irreversible inhibitor, a reversible inhibitor can dissociate from the enzyme. Reversible inhibitors include competitive inhibitors and noncompetitive inhibitors. (There are additional types of reversible inhibitors.) A competitive inhibitor is any compound that bears a structural resemblance to a particular substrate and thus competes with that substrate for binding at the active site of an enzyme. The inhibitor is not acted on by the enzyme but does prevent the substrate from approaching the active site.
The degree to which a competitive inhibitor interferes with an enzyme’s activity depends on the relative concentrations of the substrate and the inhibitor. If the inhibitor is present in relatively large quantities, it will initially block most of the active sites. But because the binding is reversible, some substrate molecules will eventually bind to the active site and be converted to product. Increasing the substrate concentration promotes displacement of the inhibitor from the active site. Competitive inhibition can be completely reversed by adding substrate so that it reaches a much higher concentration than that of the inhibitor.
Studies of competitive inhibition have provided helpful information about certain enzyme-substrate complexes and the interactions of specific groups at the active sites. As a result, pharmaceutical companies have synthesized drugs that competitively inhibit metabolic processes in bacteria and certain cancer cells. Many drugs are competitive inhibitors of specific enzymes.
A classic example of competitive inhibition is the effect of malonate on the enzyme activity of succinate dehydrogenase (Figure 9.14 "Competitive Inhibition"). Malonate and succinate are the anions of dicarboxylic acids and contain three and four carbon atoms, respectively. The malonate molecule binds to the active site because the spacing of its carboxyl groups is not greatly different from that of succinate. However, no catalytic reaction occurs because malonate does not have a CH2CH2 group to convert to CH=CH. This reaction will also be discussed in connection with the Krebs cycle and energy production. (For more information about the Krebs cycle, see Chapter 11 "Metabolic Pathways and Energy Production", Section 11.4 "Stage III of Catabolism".)
Figure 9.14 Competitive Inhibition
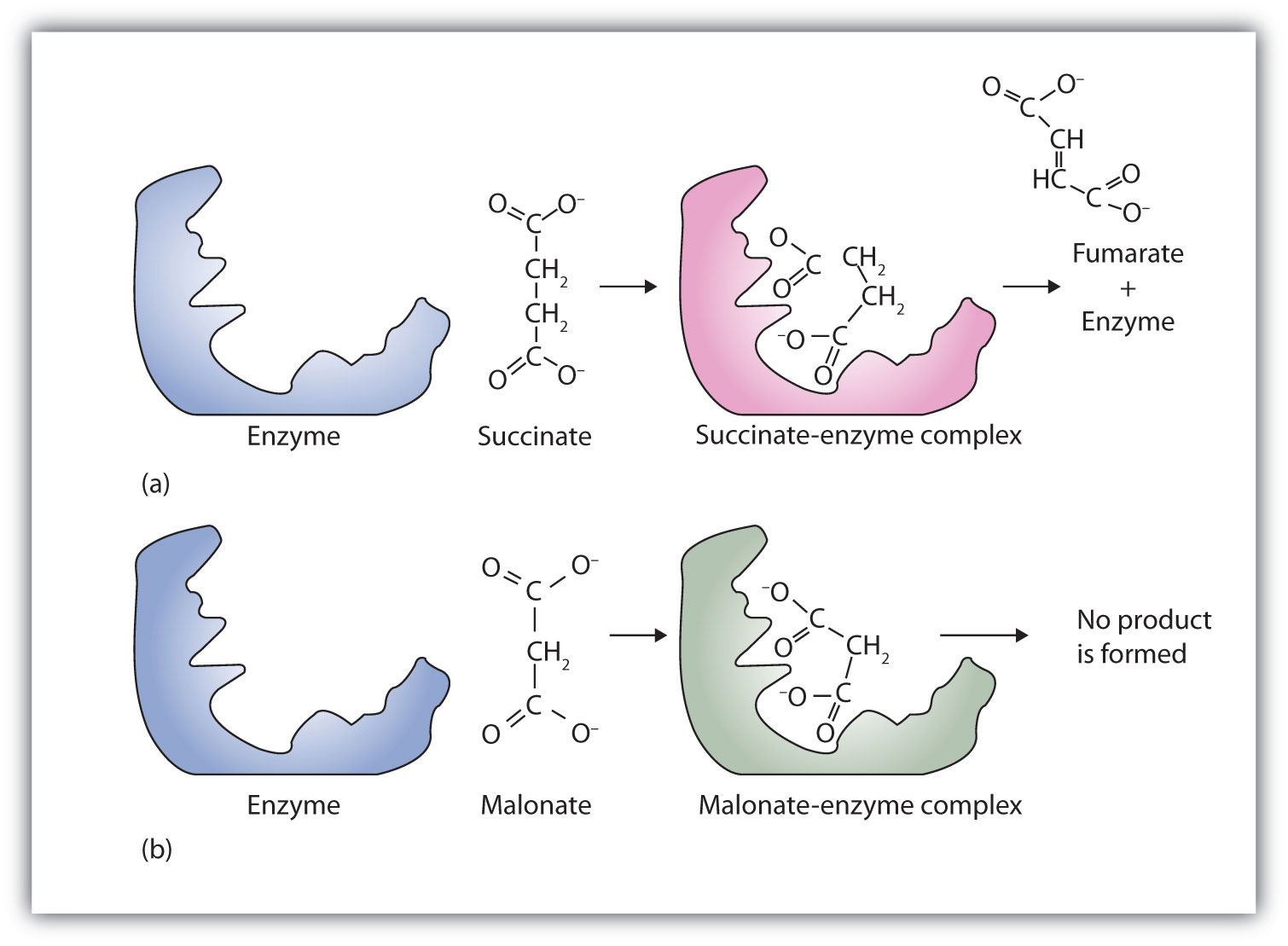
(a) Succinate binds to the enzyme succinate dehydrogenase. A dehydrogenation reaction occurs, and the product—fumarate—is released from the enzyme. (b) Malonate also binds to the active site of succinate dehydrogenase. In this case, however, no subsequent reaction occurs while malonate remains bound to the enzyme.
Chemotherapy is the strategic use of chemicals (that is, drugs) to destroy infectious microorganisms or cancer cells without causing excessive damage to the other, healthy cells of the host. From bacteria to humans, the metabolic pathways of all living organisms are quite similar, so the search for safe and effective chemotherapeutic agents is a formidable task. Many well-established chemotherapeutic drugs function by inhibiting a critical enzyme in the cells of the invading organism.
An antibiotic is a compound that kills bacteria; it may come from a natural source such as molds or be synthesized with a structure analogous to a naturally occurring antibacterial compound. Antibiotics constitute no well-defined class of chemically related substances, but many of them work by effectively inhibiting a variety of enzymes essential to bacterial growth.
Penicillin, one of the most widely used antibiotics in the world, was fortuitously discovered by Alexander Fleming in 1928, when he noticed antibacterial properties in a mold growing on a bacterial culture plate. In 1938, Ernst Chain and Howard Florey began an intensive effort to isolate penicillin from the mold and study its properties. The large quantities of penicillin needed for this research became available through development of a corn-based nutrient medium that the mold loved and through the discovery of a higher-yielding strain of mold at a United States Department of Agriculture research center near Peoria, Illinois. Even so, it was not until 1944 that large quantities of penicillin were being produced and made available for the treatment of bacterial infections.
Penicillin functions by interfering with the synthesis of cell walls of reproducing bacteria. It does so by inhibiting an enzyme—transpeptidase—that catalyzes the last step in bacterial cell-wall biosynthesis. The defective walls cause bacterial cells to burst. Human cells are not affected because they have cell membranes, not cell walls.
Several naturally occurring penicillins have been isolated. They are distinguished by different R groups connected to a common structure: a four-member cyclic amide (called a lactam ring) fused to a five-member ring. The addition of appropriate organic compounds to the culture medium leads to the production of the different kinds of penicillin.
The penicillins are effective against gram-positive bacteria (bacteria capable of being stained by Gram’s stain) and a few gram-negative bacteria (including the intestinal bacterium Escherichia coli). They are effective in the treatment of diphtheria, gonorrhea, pneumonia, syphilis, many pus infections, and certain types of boils. Penicillin G was the earliest penicillin to be used on a wide scale. However, it cannot be administered orally because it is quite unstable; the acidic pH of the stomach converts it to an inactive derivative. The major oral penicillins—penicillin V, ampicillin, and amoxicillin—on the other hand, are acid stable.
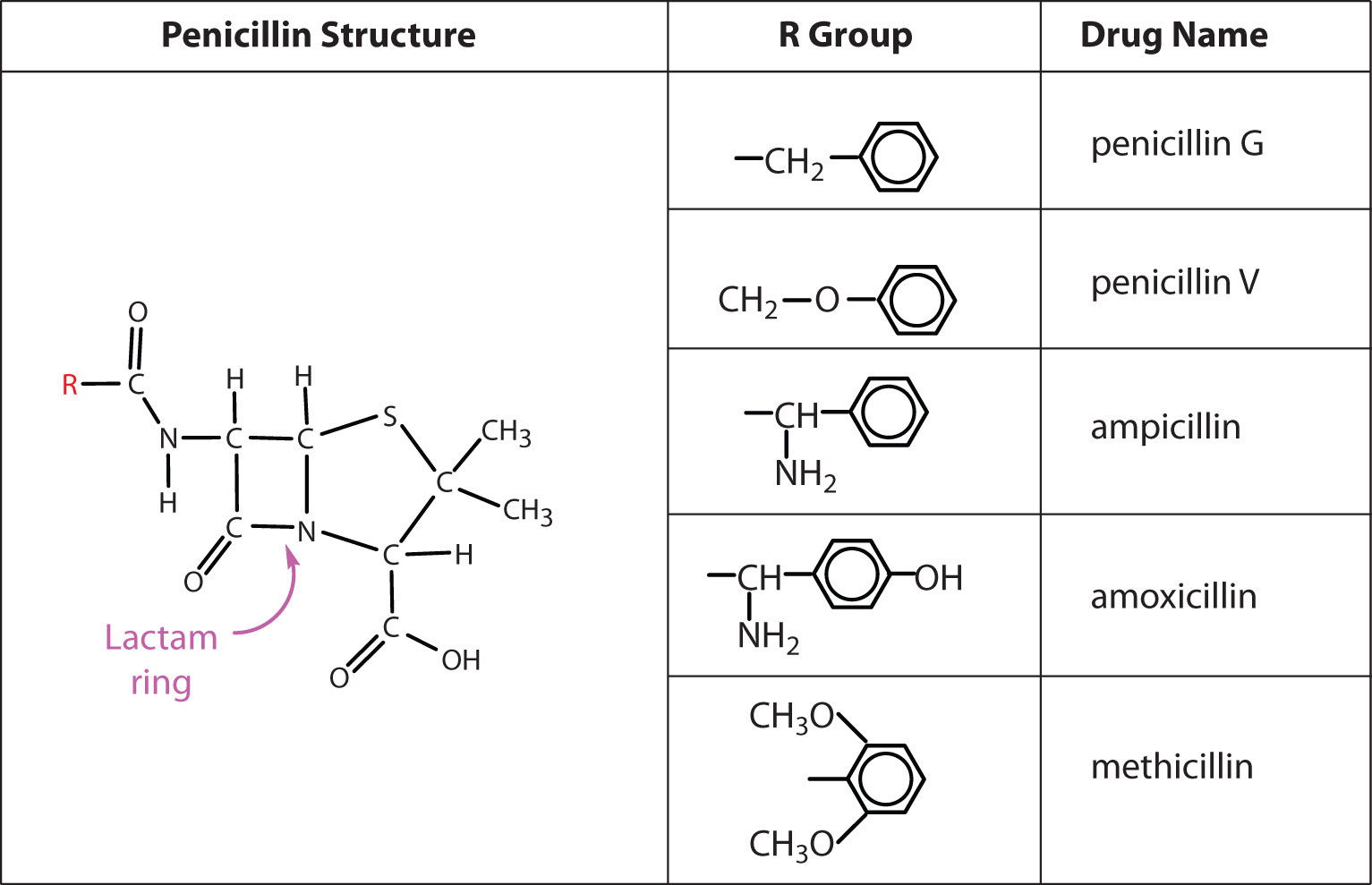
Some strains of bacteria become resistant to penicillin through a mutation that allows them to synthesize an enzyme—penicillinase—that breaks the antibiotic down (by cleavage of the amide linkage in the lactam ring). To combat these strains, scientists have synthesized penicillin analogs (such as methicillin) that are not inactivated by penicillinase.
Some people (perhaps 5% of the population) are allergic to penicillin and therefore must be treated with other antibiotics. Their allergic reaction can be so severe that a fatal coma may occur if penicillin is inadvertently administered to them. Fortunately, several other antibiotics have been discovered. Most, including aureomycin and streptomycin, are the products of microbial synthesis. Others, such as the semisynthetic penicillins and tetracyclines, are made by chemical modifications of antibiotics; and some, like chloramphenicol, are manufactured entirely by chemical synthesis. They are as effective as penicillin in destroying infectious microorganisms. Many of these antibiotics exert their effects by blocking protein synthesis in microorganisms.
Initially, antibiotics were considered miracle drugs, substantially reducing the number of deaths from blood poisoning, pneumonia, and other infectious diseases. Some seven decades ago, a person with a major infection almost always died. Today, such deaths are rare. Seven decades ago, pneumonia was a dreaded killer of people of all ages. Today, it kills only the very old or those ill from other causes. Antibiotics have indeed worked miracles in our time, but even miracle drugs have limitations. Not long after the drugs were first used, disease organisms began to develop strains resistant to them. In a race to stay ahead of resistant bacterial strains, scientists continue to seek new antibiotics. The penicillins have now been partially displaced by related compounds, such as the cephalosporins and vancomycin. Unfortunately, some strains of bacteria have already shown resistance to these antibiotics.
Some reversible inhibitors are noncompetitive. A noncompetitive inhibitor can combine with either the free enzyme or the enzyme-substrate complex because its binding site on the enzyme is distinct from the active site. Binding of this kind of inhibitor alters the three-dimensional conformation of the enzyme, changing the configuration of the active site with one of two results. Either the enzyme-substrate complex does not form at its normal rate, or, once formed, it does not yield products at the normal rate. Because the inhibitor does not structurally resemble the substrate, the addition of excess substrate does not reverse the inhibitory effect.
Feedback inhibition is a normal biochemical process that makes use of noncompetitive inhibitors to control some enzymatic activity. In this process, the final product inhibits the enzyme that catalyzes the first step in a series of reactions. Feedback inhibition is used to regulate the synthesis of many amino acids. For example, bacteria synthesize isoleucine from threonine in a series of five enzyme-catalyzed steps. As the concentration of isoleucine increases, some of it binds as a noncompetitive inhibitor to the first enzyme of the series (threonine deaminase), thus bringing about a decrease in the amount of isoleucine being formed (Figure 9.15 "Feedback Inhibition of Threonine Deaminase by Isoleucine").
Figure 9.15 Feedback Inhibition of Threonine Deaminase by Isoleucine
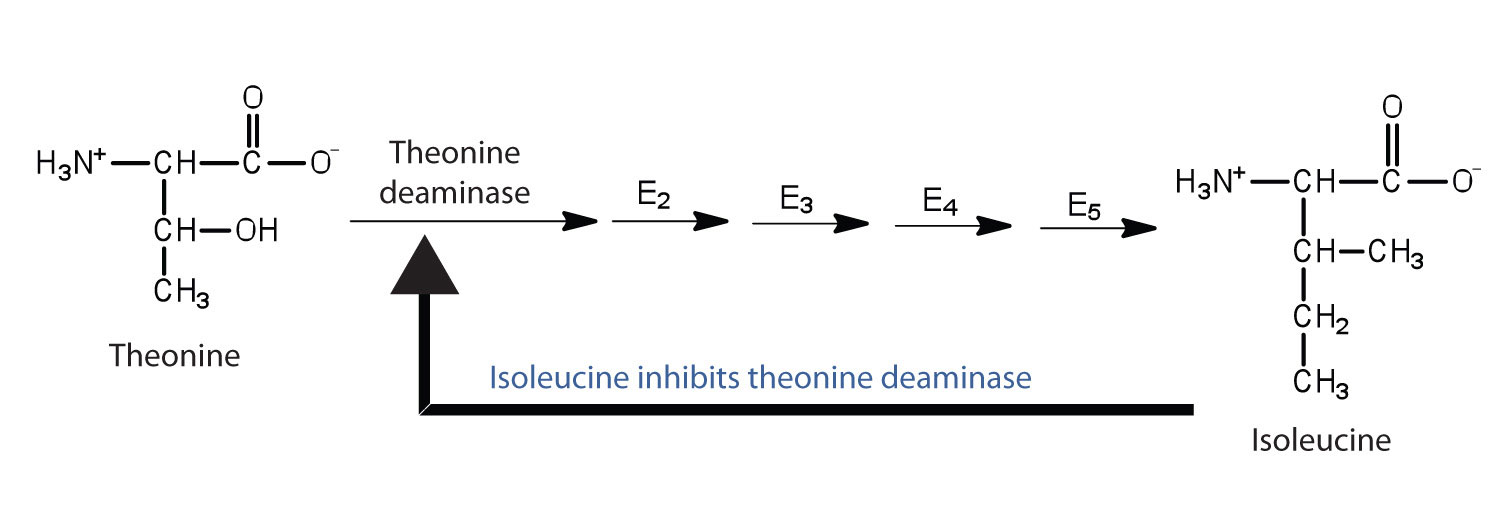
Threonine deaminase is the first enzyme in the conversion of threonine to isoleucine. Isoleucine inhibits threonine deaminase through feedback inhibition .
What are the characteristics of an irreversible inhibitor?
In what ways does a competitive inhibitor differ from a noncompetitive inhibitor?
It inactivates an enzyme by bonding covalently to a particular group at the active site.
A competitive inhibitor structurally resembles the substrate for a given enzyme and competes with the substrate for binding at the active site of the enzyme. A noncompetitive inhibitor binds at a site distinct from the active site and can bind to either the free enzyme or the enzyme-substrate complex.
What amino acid is present in the active site of all enzymes that are irreversibly inhibited by nerve gases such as DIFP?
Oxaloacetate (OOCCH2COCOO) inhibits succinate dehydrogenase. Would you expect oxaloacetate to be a competitive or noncompetitive inhibitor? Explain.
serine
Many enzymes are simple proteins consisting entirely of one or more amino acid chains. Other enzymes contain a nonprotein component called a cofactor that is necessary for the enzyme’s proper functioning. There are two types of cofactors: inorganic ions [e.g., zinc or Cu(I) ions] and organic molecules known as coenzymes. Most coenzymes are vitamins or are derived from vitamins.
Vitamins are organic compounds that are essential in very small (trace) amounts for the maintenance of normal metabolism. They generally cannot be synthesized at adequate levels by the body and must be obtained from the diet. The absence or shortage of a vitamin may result in a vitamin-deficiency disease. In the first half of the 20th century, a major focus of biochemistry was the identification, isolation, and characterization of vitamins.
Despite accumulating evidence that people needed more than just carbohydrates, fats, and proteins in their diets for normal growth and health, it was not until the early 1900s that research established the need for trace nutrients in the diet.
Because organisms differ in their synthetic abilities, a substance that is a vitamin for one species may not be so for another. Over the past 100 years, scientists have identified and isolated 13 vitamins required in the human diet and have divided them into two broad categories: the fat-soluble vitamins, which include vitamins A, D, E, and K, and the water-soluble vitamins, which are the B complex vitamins and vitamin C. All fat-soluble vitamins contain a high proportion of hydrocarbon structural components. There are one or two oxygen atoms present, but the compounds as a whole are nonpolar. In contrast, water-soluble vitamins contain large numbers of electronegative oxygen and nitrogen atoms, which can engage in hydrogen bonding with water. Most water-soluble vitamins act as coenzymes or are required for the synthesis of coenzymes. The fat-soluble vitamins are important for a variety of physiological functions. The key vitamins and their functions are found in Table 9.5 "Fat-Soluble Vitamins and Physiological Functions" and Table 9.6 "Water-Soluble Vitamins and Physiological Functions".
Table 9.5 Fat-Soluble Vitamins and Physiological Functions
| Vitamin | Physiological Function | Effect of Deficiency |
|---|---|---|
| vitamin A (retinol) | formation of vision pigments; differentiation of epithelial cell | night blindness; continued deficiency leads to total blindness |
| vitamin D (cholecalciferol) | increases the body’s ability to absorb calcium and phosphorus | osteomalacia (softening of the bones); known as rickets in children |
| vitamin E (tocopherol) | fat-soluble antioxidant | damage to cell membranes |
| vitamin K (phylloquinone) | formation of prothrombin, a key enzyme in the blood-clotting process | increases the time required for blood to clot |
Table 9.6 Water-Soluble Vitamins and Physiological Functions
| Vitamin | Coenzyme | Coenzyme Function | Deficiency Disease |
|---|---|---|---|
| vitamin B1 (thiamine) | thiamine pyrophosphate | decarboxylation reactions | beri-beri |
| vitamin B2 (riboflavin) | flavin mononucleotide or flavin adenine dinucleotide | oxidation-reduction reactions involving two hydrogen atoms | — |
| vitamin B3 (niacin) | nicotinamide adenine dinucleotide or nicotinamide adenine dinucleotide phosphate | oxidation-reduction reactions involving the hydride ion (H−) | pellagra |
| vitamin B6 (pyridoxine) | pyridoxal phosphate | variety of reactions including the transfer of amino groups | — |
| vitamin B12 (cyanocobalamin) | methylcobalamin or deoxyadenoxylcobalamin | intramolecular rearrangement reactions | pernicious anemia |
| biotin | biotin | carboxylation reactions | — |
| folic acid | tetrahydrofolate | carrier of one-carbon units such as the formyl group | anemia |
| pantothenic Acid | coenzyme A | carrier of acyl groups | — |
| vitamin C (ascorbic acid) | none | antioxidant; formation of collagen, a protein found in tendons, ligaments, and bone | scurvy |
Vitamins C and E, as well as the provitamin β-carotene can act as antioxidants in the body. Antioxidants prevent damage from free radicals, which are molecules that are highly reactive because they have unpaired electrons. Free radicals are formed not only through metabolic reactions involving oxygen but also by such environmental factors as radiation and pollution.
β-carotene is known as a provitamin because it can be converted to vitamin A in the body.
Free radicals react most commonly react with lipoproteins and unsaturated fatty acids in cell membranes, removing an electron from those molecules and thus generating a new free radical. The process becomes a chain reaction that finally leads to the oxidative degradation of the affected compounds. Antioxidants react with free radicals to stop these chain reactions by forming a more stable molecule or, in the case of vitamin E, a free radical that is much less reactive. (Vitamin E is converted back to its original form through interaction with vitamin C.)
What is the difference between a cofactor and a coenzyme?
How are vitamins related to coenzymes?
A coenzyme is one type of cofactor. Coenzymes are organic molecules required by some enzymes for activity. A cofactor can be either a coenzyme or an inorganic ion.
Coenzymes are synthesized from vitamins.
1. Identify each vitamin as water soluble or fat soluble.
a. vitamin D
b. vitamin C
c. vitamin B12
2. Identify each vitamin as water soluble or fat soluble.
a. niacin
b. cholecalciferol
c. biotin
3. What vitamin is needed to form each coenzyme?
a. pyridoxal phosphate
b. flavin adenine dinucleotide
c. coenzyme A
d. nicotinamide adenine dinucleotide
4. What coenzyme is formed from each vitamin?
a. niacin
b. thiamine
c. cyanocobalamin
d. pantothenic acid
5. What is the function of each vitamin or coenzyme?
a. flavin adenine dinucleotide
b. vitamin A
c. biotin
6. What is the function of each vitamin or coenzyme?
a. vitamin K
b. pyridoxal phosphate
c. tetrahydrofolate
1.
a. fat soluble
b. water soluble
c. water soluble
3.
a. vitamin B6 or pyridoxine
b. vitamin B2 or riboflavin
c. pantothenic acid
d. vitamin B3 or niacin
5.
a. needed by enzymes that catalyze oxidation-reduction reactions in which two hydrogen atoms are transferred
b. needed for the formation of vision pigments
c. needed by enzymes that catalyze carboxylation reactions
To ensure that you understand the material in this chapter, you should review the meanings of the bold terms in the following summary and ask yourself how they relate to the topics in the chapter.
A protein is a large biological polymer synthesized from amino acids, which are carboxylic acids containing an α-amino group. Proteins have a variety of important roles in living organisms, yet they are made from the same 20 L-amino acids. About half of these amino acids, the essential amino acids, cannot be synthesized by the human body and must be obtained from the diet. In the solid state and in neutral solutions, amino acids exist as zwitterions, species that are charged but electrically neutral. In this form, they behave much like inorganic salts. Each amino acid belongs to one of four classes depending on the characteristics of its R group or amino acid side chain: nonpolar, polar but neutral, positively charged, and negatively charged. Depending on the conditions, amino acids can act as either acids or bases, which means that proteins act as buffers. The pH at which an amino acid exists as the zwitterion is called the isoelectric point (pI).
The amino acids in a protein are linked together by peptide bonds. Protein chains containing 10 or fewer amino acids are usually referred to as peptides, with a prefix such as di- or tri- indicating the number of amino acids. Chains containing more than 50 amino acid units are referred to as proteins or polypeptides. Proteins are classified globular or fibrous, depending on their structure and resulting solubility in water. Globular proteins are nearly spherical and are soluble in water; fibrous proteins have elongated or fibrous structures and are not soluble in water.
Protein molecules can have as many as four levels of structure. The primary structure is the sequence of amino acids in the chain. The secondary structure is the arrangement of adjacent atoms in the peptide chain; the most common arrangements are α-helices or β-pleated sheets. The tertiary structure is the overall three-dimensional shape of the molecule that results from the way the chain bends and folds in on itself. Proteins that consist of more than one chain have quaternary structure, which is the way the multiple chains are packed together.
Four types of intramolecular and intermolecular forces contribute to secondary, tertiary, and quaternary structure: (1) hydrogen bonding between an oxygen or a nitrogen atom and a hydrogen atom bound to an oxygen atom or a nitrogen atom, either on the same chain or on a neighboring chain; (2) ionic bonding between one positively charged side chain and one negatively charged side chain; (3) disulfide linkages between cysteine units; and (4) dispersion forces between nonpolar side chains.
Because of their complexity, protein molecules are delicate and easy to disrupt. A denatured protein is one whose conformation has been changed, in a process called denaturation, so that it can no longer do its physiological job. A variety of conditions, such as heat, ultraviolet radiation, the addition of organic compounds, or changes in pH can denature a protein.
An enzyme is an organic catalyst produced by a living cell. Enzymes are such powerful catalysts that the reactions they promote occur rapidly at body temperature. Without the help of enzymes, these reactions would require high temperatures and long reaction times.
The molecule or molecules on which an enzyme acts are called its substrates. An enzyme has an active site where its substrate or substrates bind to form an enzyme-substrate complex. The reaction occurs, and product is released:
E + S → E–S → E + PThe original lock-and-key model of enzyme and substrate binding pictured a rigid enzyme of unchanging configuration binding to the appropriate substrate. The newer induced-fit model describes the enzyme active site as changing its conformation after binding to the substrate.
Most enzymes have maximal activity in a narrow pH range centered on an optimum pH. In this pH range, the enzyme is correctly folded, and catalytic groups in the active site have the correct charge (positive, negative, or neutral). For most enzymes, the optimum pH is between 6 and 8.
Substances that interfere with enzyme function are called inhibitors. An irreversible inhibitor inactivates enzymes by forming covalent bonds to the enzyme, while a reversible inhibitor inactivates an enzyme by a weaker, noncovalent interaction that is easier to disrupt. A competitive inhibitor is a reversible inhibitor that is structurally similar to the substrate and binds to the active site. When the inhibitor is bound, the substrate is blocked from the active site and no reaction occurs. Because the binding of such an inhibitor is reversible, a high substrate concentration will overcome the inhibition because it increases the likelihood of the substrate binding. A noncompetitive inhibitor binds reversibly at a site distinct from the active site. Thus, it can bind to either the enzyme or the enzyme-substrate complex. The inhibitor changes the conformation of the active site so that the enzyme cannot function properly. Noncompetitive inhibitors are important in feedback inhibition, in which the amount of product produced by a series of reactions is carefully controlled. The final product in a series of reactions acts as a noncompetitive inhibitor of the initial enzyme.
Simple enzymes consist entirely of one or more amino acid chains. Complex enzymes are composed of one or more amino acid chains joined to cofactors—inorganic ions or organic coenzymes. Vitamins are organic compounds that are essential in very small amounts for the maintenance of normal metabolism and generally cannot be synthesized at adequate levels by the body. Vitamins are divided into two broad categories: fat-soluble vitamins and water-soluble vitamins. Many of the water-soluble vitamins are used for the synthesis of coenzymes.
1. Draw the structure of the amino acid γ-aminobutyric acid (GABA). Would you expect to find GABA in the amino acid sequence of a protein? Explain.
2. Draw the structure of the amino acid homocysteine (R group = CH2CH2SH). Would you expect to find homocysteine in the amino acid sequence of a protein? Justify your answer.
3. Write equations to show how leucine can act as a buffer (that is, how it can neutralize added acid or base).
4. Write equations to show how isoleucine can act as a buffer (that is, how it can neutralize added acid or base).
5. Glutathione (γ-glutamylcysteinylglycine) is a tripeptide found in all cells of higher animals. It contains glutamic acid joined in an unusual peptide linkage involving the carboxyl group of the R group (known as γ-carboxyl group), rather than the usual carboxyl group (the α-carboxyl group). Draw the structure of glutathione.
6. Draw the structure of the pentapeptide whose sequence is arg-his-gly-leu-asp. Identify which of the amino acids have R groups that can donate or gain hydrogen ions.
7. Bradykinin is a peptide hormone composed of nine amino acids that lowers blood pressure. Its primary structure is arg-pro-pro-gly-phe-ser-pro-phe-arg. Would you expect bradykinin to be positively charged, negatively charged, or neutral at a pH of 6.0? Justify your answer.
8. One of the neurotransmitters involved in pain sensation is a peptide called substance P, which is composed of 11 amino acids and is released by nerve-cell terminals in response to pain. Its primary structure is arg-pro-lys-pro-gln-gln-phe-phe-gly-leu-met. Would you expect this peptide to be positively charged, negatively charged, or neutral at a pH of 6.0? Justify your answer.
9. Carbohydrates are incorporated into glycoproteins. Would you expect the incorporation of sugar units to increase or decrease the solubility of a protein? Justify your answer.
10. Some proteins have phosphate groups attached through an ester linkage to the OH groups of serine, threonine, or tyrosine residues to form phosphoproteins. Would you expect the incorporation of a phosphate group to increase or decrease the solubility of a protein? Justify your answer.
11. Refer to Table 9.2 "Classes of Enzymes" and determine how each enzyme would be classified.
a. the enzyme that catalyzes the conversion of ethanol to acetaldehyde
b. the enzyme that catalyzes the breakdown of glucose 6-phosphate to glucose and inorganic phosphate ion (water is also a reactant in this reaction)
12. Refer to Table 9.2 "Classes of Enzymes" and determine how each enzyme would be classified.
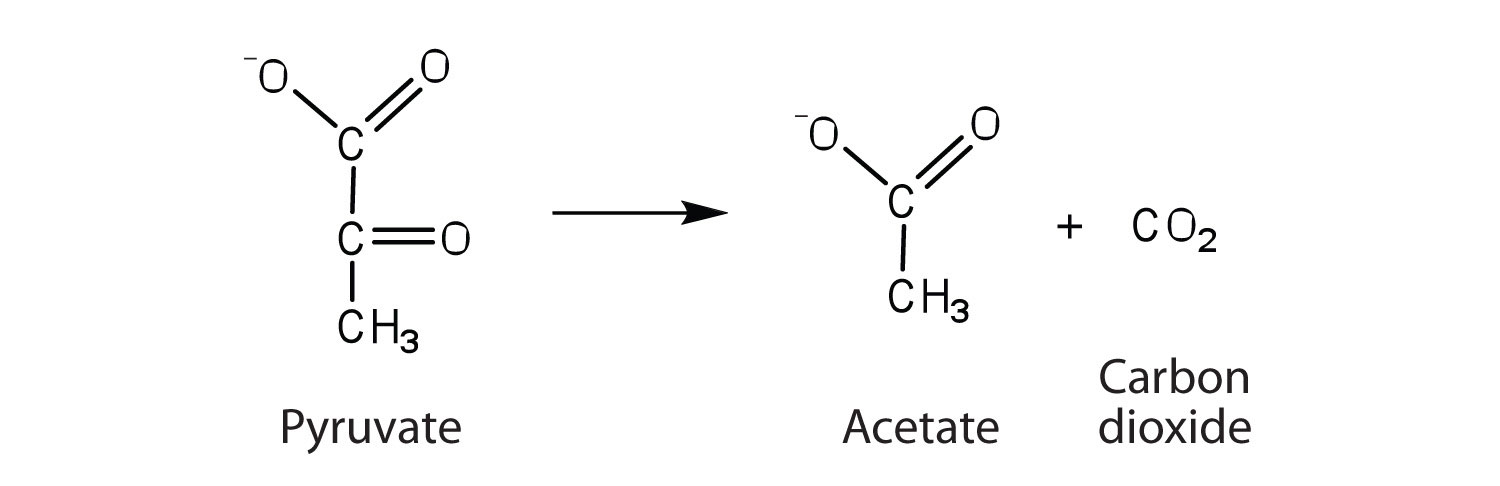
a. the enzyme that catalyzes the removal of a carboxyl group from pyruvate to form acetate
b. the enzyme that catalyzes the rearrangement of 3-phosphoglycerate to form 2-phosphoglycerate
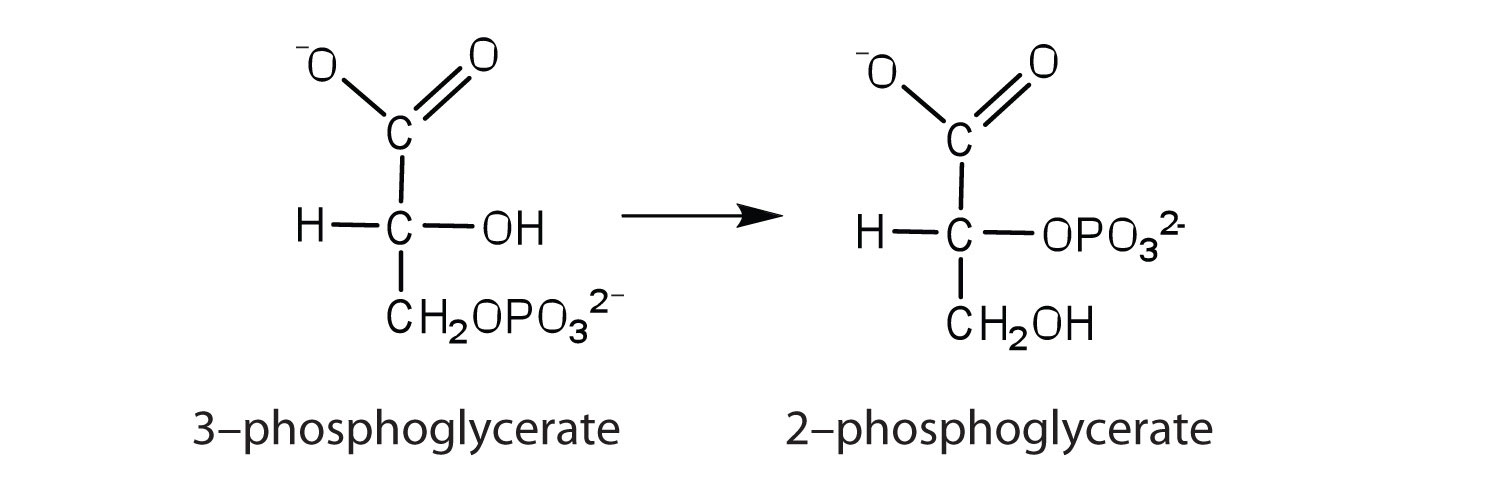
13. The enzyme lysozyme has an aspartic acid residue in the active site. In acidic solution, the enzyme is inactive, but activity increases as the pH rises to around 6. Explain why.
14. The enzyme lysozyme has a glutamic acid residue in the active site. At neutral pH (6–7), the enzyme is active, but activity decreases as the pH rises. Explain why.
15. The activity of a purified enzyme is measured at a substrate concentration of 1.0 μM and found to convert 49 μmol of substrate to product in 1 min. The activity is measured at 2.0 μM substrate and found to convert 98 μmol of substrate to product/minute.
a. When the substrate concentration is 100 μM, how much substrate would you predict is converted to product in 1 min? What if the substrate concentration were increased to 1,000 μM (1.0 mM)?
b. The activities actually measured are 676 μmol product formed/minute at a substrate concentration of 100 μM and 698 μmol product formed/minute at 1,000 μM (1.0 mM) substrate. Is there any discrepancy between these values and those you predicted in Exercise 15a? Explain.
16. A patient has a fever of 39°C. Would you expect the activity of enzymes in the body to increase or decrease relative to their activity at normal body temperature (37°C)?
17. Using your knowledge of factors that influence enzyme activity, describe what happens when milk is pasteurized.
1. This amino acid would not be found in proteins because it is not an α-amino acid.
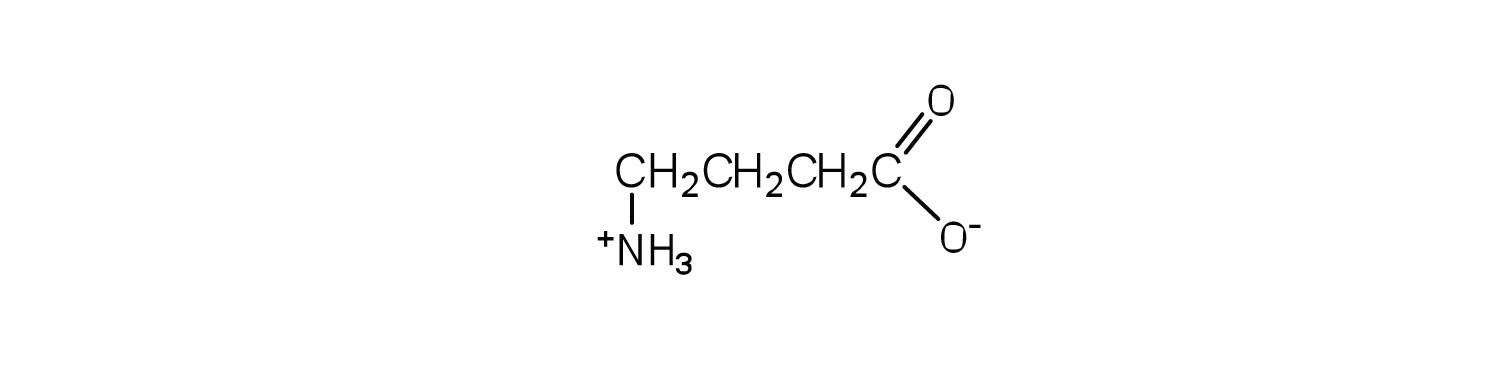
3.
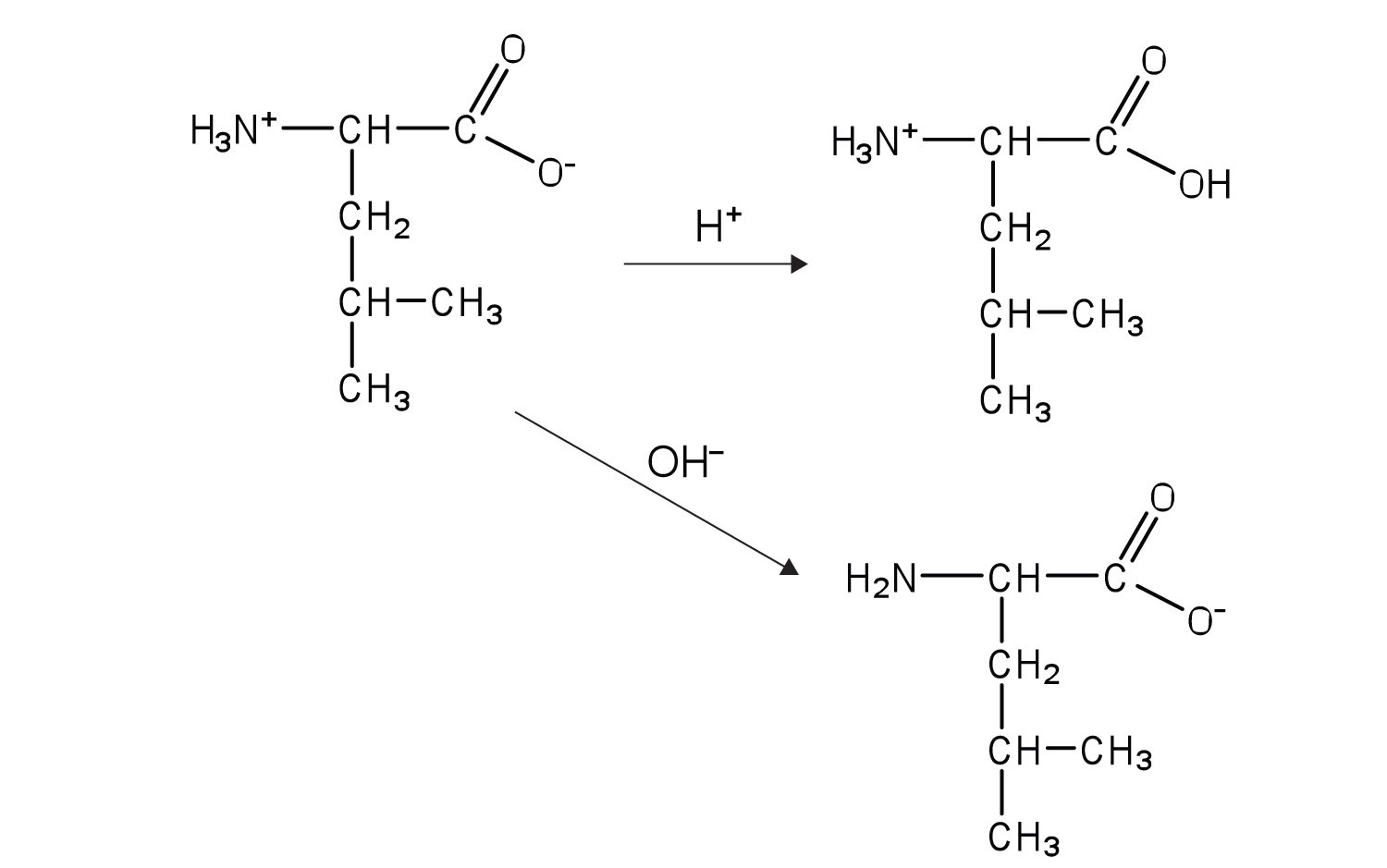
5.
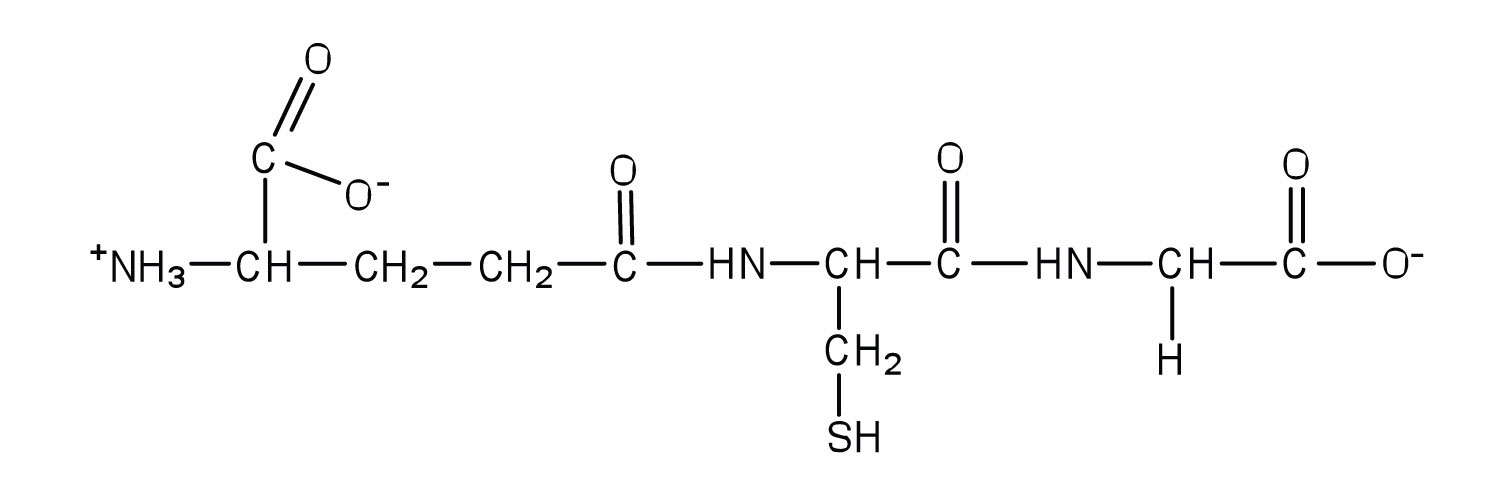
7. Bradykinin would be positively charged; all of the amino acids, except for arginine, have R groups that do not become either positively or negatively charged. The two arginines are R groups that are positively charged at neutral pH, so the peptide would have an overall positive charge.
9. Carbohydrates have many OH groups attached, which can engage in hydrogen bonding with water, which increases the solubility of the proteins.
11.
a. oxidoreductase
b. hydrolase
13. The enzyme is active when the carboxyl group in the R group of aspartic acid does not have the hydrogen attached (forming COO–); the hydrogen is removed when the pH of the solution is around pH 6 or higher.
15.
a. at 100 μM, you would predict that the rate would increase 100 times to 4,900 μmol of substrate to product in 1 min; at 1.0 mM, you would predict an increase to 49,000 μmol of substrate to product in 1 min.
b. There is a great discrepancy between the predicted rates and actual rates; this occurs because the enzyme becomes saturated with substrate, preventing a further increase in the rate of the reaction (the reaction is no longer linear with respect to substrate concentration because it is at very low concentrations).
17. When milk is pasteurized, it is heated to high temperatures. These high temperatures denature the proteins in bacteria, so they cannot carry out needed functions to grow and multiply.
Library Info and Research Help | reflibrarian@hostos.cuny.edu (718) 518-4215
Loans or Fines | circ@hostos.cuny.edu (718) 518-4222
475 Grand Concourse (A Building), Room 308, Bronx, NY 10451
